Chromatin regulatory mechanisms in pluripotency
- PMID: 20624054
- PMCID: PMC3085914
- DOI: 10.1146/annurev-cellbio-051809-102012
Chromatin regulatory mechanisms in pluripotency
Abstract
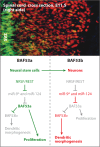
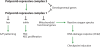
Stem cells of all types are characterized by a stable, heritable state permissive of multiple developmental pathways. The past five years have seen remarkable advances in understanding these heritable states and the ways that they are initiated or terminated. Transcription factors that bind directly to DNA and have sufficiency roles have been most easy to investigate and, perhaps for this reason, are most solidly implicated in pluripotency. In addition, large complexes of ATP-dependent chromatin-remodeling and histone-modification enzymes that have specialized functions have also been implicated by genetic studies in initiating and/or maintaining pluripotency or multipotency. Several of these ATP-dependent remodeling complexes play non-redundant roles, and the esBAF complex facilitates reprogramming of induced pluripotent stem cells. The recent finding that virtually all histone modifications can be rapidly reversed and are often highly dynamic has raised new questions about how histone modifications come to play a role in the steady state of pluripotency. Another surprise from genetic studies has been the frequency with which the global effects of mutations in chromatin regulators can be largely reversed by a single target gene. These genetic studies help define the arena for future mechanistic studies that might be helpful to harness pluripotency for therapeutic goals.
Figures


References
-
- Agger K, Cloos PA, Christensen J, Pasini D, Rose S, et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature. 2007;449(7163):731–34. - PubMed
-
- Azuara V, Perry P, Sauer S, Spivakov M, Jorgensen HF, et al. Chromatin signatures of pluripotent cell lines. Nat. Cell Biol. 2006;8(5):532–38. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

