Genome-wide real-time PCR for West Nile virus reduces the false-negative rate and facilitates new strain discovery
- PMID: 20637239
- PMCID: PMC3189403
- DOI: 10.1016/j.jviromet.2010.07.005
Genome-wide real-time PCR for West Nile virus reduces the false-negative rate and facilitates new strain discovery
Abstract
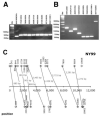
West Nile virus (WNV) causes significant morbidity and mortality worldwide. Transplant and transfusion recipients as well as the elderly are particularly at risk. WNV shows strain variation from season to season and from locale to locale. This poses a significant problem for diagnosis. Most assays use a single primer pair to detect WNV by QPCR, and can fail to detect novel stains. To overcome this limitation, a genome-wide, multiple primer-based real-time QPCR assay was developed for WNV. The same assay can be used for quantitation, viral variant discovery as well as for amplification of the entire viral genome using a single annealing temperature. It improves upon routine diagnosis as well as facilitates laboratory investigations of the pathology of WNV.
Copyright (c) 2010 Elsevier B.V. All rights reserved.
Figures


Similar articles
-
Real time PCR assay for detection of all known lineages of West Nile virus.J Virol Methods. 2016 Oct;236:266-270. doi: 10.1016/j.jviromet.2016.07.026. Epub 2016 Jul 30. J Virol Methods. 2016. PMID: 27481597
-
A novel quantitative multiplex real-time RT-PCR for the simultaneous detection and differentiation of West Nile virus lineages 1 and 2, and of Usutu virus.J Virol Methods. 2013 May;189(2):321-7. doi: 10.1016/j.jviromet.2013.02.019. Epub 2013 Mar 7. J Virol Methods. 2013. PMID: 23499258
-
Highly sensitive TaqMan RT-PCR assay for detection and quantification of both lineages of West Nile virus RNA.J Clin Virol. 2006 Jul;36(3):177-82. doi: 10.1016/j.jcv.2006.02.008. Epub 2006 May 3. J Clin Virol. 2006. PMID: 16675298
-
Application of West Nile virus diagnostic techniques.Expert Rev Anti Infect Ther. 2013 Aug;11(8):793-803. doi: 10.1586/14787210.2013.814824. Expert Rev Anti Infect Ther. 2013. PMID: 23977935 Review.
-
Genetic systems of West Nile virus and their potential applications.Curr Opin Investig Drugs. 2003 Aug;4(8):959-65. Curr Opin Investig Drugs. 2003. PMID: 14508880 Review.
Cited by
-
Parasite Tolerance and Host Competence in Avian Host Defense to West Nile Virus.Ecohealth. 2018 Jun;15(2):360-371. doi: 10.1007/s10393-018-1332-7. Epub 2018 Mar 22. Ecohealth. 2018. PMID: 29569179
-
Recent progress in West Nile virus diagnosis and vaccination.Vet Res. 2012 Mar 1;43(1):16. doi: 10.1186/1297-9716-43-16. Vet Res. 2012. PMID: 22380523 Free PMC article. Review.
-
Evolutionary dynamics and molecular epidemiology of West Nile virus in New York State: 1999-2015.Virus Evol. 2019 Jul 21;5(2):vez020. doi: 10.1093/ve/vez020. eCollection 2019 Jul. Virus Evol. 2019. PMID: 31341640 Free PMC article.
-
Diagnosis of west nile virus human infections: overview and proposal of diagnostic protocols considering the results of external quality assessment studies.Viruses. 2013 Sep 25;5(10):2329-48. doi: 10.3390/v5102329. Viruses. 2013. PMID: 24072061 Free PMC article. Review.
-
Development of a multi-target TaqMan assay to detect eastern equine encephalitis virus variants in mosquitoes.Vector Borne Zoonotic Dis. 2012 Oct;12(10):872-6. doi: 10.1089/vbz.2012.1008. Epub 2012 Jul 26. Vector Borne Zoonotic Dis. 2012. PMID: 22835151 Free PMC article.
References
-
- Armstrong WS, Bashour CA, Smedira NG, Heupler FA, Hoeltge GA, Mawhorter SD, Sudheendra V, Gordon SM. A case of fatal West Nile virus meningoencephalitis associated with receipt of blood transfusions after open heart surgery. Ann Thorac Surg. 2003;76:605–7. - PubMed
-
- Beasley DW, Li L, Suderman MT, Barrett AD. Mouse neuroinvasive phenotype of West Nile virus strains varies depending upon virus genotype. Virology. 2002;296:17–23. - PubMed
-
- Busch MP, Caglioti S, Robertson EF, McAuley JD, Tobler LH, Kamel H, Linnen JM, Shyamala V, Tomasulo P, Kleinman SH. Screening the blood supply for West Nile virus RNA by nucleic acid amplification testing. N Engl J Med. 2005;353:460–7. - PubMed
-
- Cantor CR, Schimmel PR. Biophysical Chemistry: The Behavior of Biological Macromolecules. Freeman; New York: 1980.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

