Analysis of infectious virus clones from two HIV-1 superinfection cases suggests that the primary strains have lower fitness
- PMID: 20646276
- PMCID: PMC2918528
- DOI: 10.1186/1742-4690-7-60
Analysis of infectious virus clones from two HIV-1 superinfection cases suggests that the primary strains have lower fitness
Abstract
Background: Two HIV-1 positive patients, L and P, participating in the Amsterdam Cohort studies acquired an HIV-1 superinfection within half a year from their primary HIV-1 infection (Jurriaans et al., JAIDS 2008, 47:69-73). The aim of this study was to compare the replicative fitness of the primary and superinfecting HIV-1 strains of both patients. The use of isolate-specific primer sets indicated that the primary and secondary strains co-exist in plasma at all time points after the moment of superinfection.
Results: Biological HIV-1 clones were derived from peripheral blood CD4 + T cells at different time point, and identified as the primary or secondary virus through sequence analysis. Replication competition assays were performed with selected virus pairs in PHA/IL-2 activated peripheral blood mononuclear cells (PBMC's) and analyzed with the Heteroduplex Tracking Assay (HTA) and isolate-specific PCR amplification. In both cases, we found a replicative advantage of the secondary HIV-1 strain over the primary virus. Full-length HIV-1 genomes were sequenced to find possible explanations for the difference in replication capacity. Mutations that could negatively affect viral replication were identified in the primary infecting strains. In patient L, the primary strain has two insertions in the LTR promoter, combined with a mutation in the tat gene that has been associated with decreased replication capacity. The primary HIV-1 strain isolated from patient P has two mutations in the LTR that have been associated with a reduced replication rate. In a luciferase assay, only the LTR from the primary virus of patient P had lower transcriptional activity compared with the superinfecting virus.
Conclusions: These preliminary findings suggest the interesting scenario that superinfection occurs preferentially in patients infected with a relatively attenuated HIV-1 isolate.
Figures






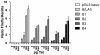
References
-
- Quinones-Mateu ME, Arts EJ. Virus fitness: concept, quantification, and application to HIV population dynamics. Curr Top Microbiol Immunol. 2006;299:83–140. full_text. - PubMed
-
- Quinones-Mateu ME, Ball SC, Marozsan AJ, Torre VS, Albright JL, Vanham G, van Der GG, Colebunders RL, Arts EJ. A dual infection/competition assay shows a correlation between ex vivo human immunodeficiency virus type 1 fitness and disease progression. J Virol. 2000;74:9222–9233. doi: 10.1128/JVI.74.19.9222-9233.2000. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials

