Mechanistic similarity and diversity among the guanidine-modifying members of the pentein superfamily
- PMID: 20654741
- PMCID: PMC4104755
- DOI: 10.1016/j.bbapap.2010.07.016
Mechanistic similarity and diversity among the guanidine-modifying members of the pentein superfamily
Abstract
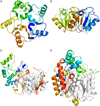
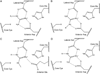
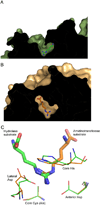
The pentein superfamily is a mechanistically diverse superfamily encompassing both noncatalytic proteins and enzymes that catalyze hydrolase, dihydrolase and amidinotransfer reactions on guanidine substrates. Despite generally low sequence identity, they possess a conserved structural fold and display common mechanistic themes in catalysis. The structurally characterized catalytic penteins possess a conserved core of residues that include a Cys, His and two polar, guanidine-binding residues. All known catalytic penteins use the core Cys to attack the substrate's guanidine moiety to form a covalent thiouronium adduct and all cleave one or more of the guanidine C--N bonds. The mechanistic information compiled to date supports the hypothesis that this superfamily may have evolved divergently from a catalytically promiscuous ancestor.
Copyright © 2010. Published by Elsevier B.V.
Figures





References
-
- Groft CM, Beckmann R, Sali A, Burley SK. Crystal structures of ribosome anti-association factor IF6. Nature Structural & Molecular Biology. 2000 Dec;7:1156–1164. - PubMed
-
- Paoli M. An elusive propeller-like fold. Nature Structural Biology. 2001 Sep;8:744–745. - PubMed
-
- Teichmann SA, Murzin AG, Chothia C, Determination of protein function. evolution and interactions by structural genomics. Current Opinion in Structural Biology. 2001 Jun;11:354–363. - PubMed
-
- Hartzoulakis B, Rossiter S, Gill H, O’Hara B, Steinke E, Gane PJ, Hurtado-Guerrero R, Leiper JM, Vallance P, Rust JM, Selwood DL. Discovery of inhibitors of the pentein superfamily protein dimethylarginine dimethylaminohydrolase (DDAH) by virtual screening and hit analysis. Bioorganic & Medicinal Chemistry Letters. 2007 Jul;17:3953–3956. - PubMed
-
- Groft CM, Beckmann R, Sali A, Burley SK. Crystal structures of ribosome anti-association factor IF6. Nature Structural Biology. 2000 Dec;7:1156–1164. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

