Broadly sampled multigene analyses yield a well-resolved eukaryotic tree of life
- PMID: 20656852
- PMCID: PMC2950834
- DOI: 10.1093/sysbio/syq037
Broadly sampled multigene analyses yield a well-resolved eukaryotic tree of life
Abstract
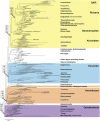
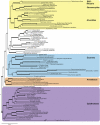
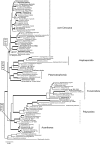
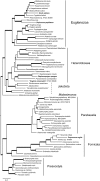
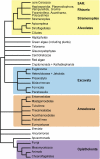
An accurate reconstruction of the eukaryotic tree of life is essential to identify the innovations underlying the diversity of microbial and macroscopic (e.g., plants and animals) eukaryotes. Previous work has divided eukaryotic diversity into a small number of high-level "supergroups," many of which receive strong support in phylogenomic analyses. However, the abundance of data in phylogenomic analyses can lead to highly supported but incorrect relationships due to systematic phylogenetic error. Furthermore, the paucity of major eukaryotic lineages (19 or fewer) included in these genomic studies may exaggerate systematic error and reduce power to evaluate hypotheses. Here, we use a taxon-rich strategy to assess eukaryotic relationships. We show that analyses emphasizing broad taxonomic sampling (up to 451 taxa representing 72 major lineages) combined with a moderate number of genes yield a well-resolved eukaryotic tree of life. The consistency across analyses with varying numbers of taxa (88-451) and levels of missing data (17-69%) supports the accuracy of the resulting topologies. The resulting stable topology emerges without the removal of rapidly evolving genes or taxa, a practice common to phylogenomic analyses. Several major groups are stable and strongly supported in these analyses (e.g., SAR, Rhizaria, Excavata), whereas the proposed supergroup "Chromalveolata" is rejected. Furthermore, extensive instability among photosynthetic lineages suggests the presence of systematic biases including endosymbiotic gene transfer from symbiont (nucleus or plastid) to host. Our analyses demonstrate that stable topologies of ancient evolutionary relationships can be achieved with broad taxonomic sampling and a moderate number of genes. Finally, taxon-rich analyses such as presented here provide a method for testing the accuracy of relationships that receive high bootstrap support (BS) in phylogenomic analyses and enable placement of the multitude of lineages that lack genome scale data.
Figures
Similar articles
-
Taxon-rich phylogenomic analyses resolve the eukaryotic tree of life and reveal the power of subsampling by sites.Syst Biol. 2015 May;64(3):406-15. doi: 10.1093/sysbio/syu126. Epub 2014 Dec 23. Syst Biol. 2015. PMID: 25540455
-
Broadly sampled multigene trees of eukaryotes.BMC Evol Biol. 2008 Jan 18;8:14. doi: 10.1186/1471-2148-8-14. BMC Evol Biol. 2008. PMID: 18205932 Free PMC article.
-
Phylogenomic analysis supports the monophyly of cryptophytes and haptophytes and the association of rhizaria with chromalveolates.Mol Biol Evol. 2007 Aug;24(8):1702-13. doi: 10.1093/molbev/msm089. Epub 2007 May 7. Mol Biol Evol. 2007. PMID: 17488740
-
The phagotrophic origin of eukaryotes and phylogenetic classification of Protozoa.Int J Syst Evol Microbiol. 2002 Mar;52(Pt 2):297-354. doi: 10.1099/00207713-52-2-297. Int J Syst Evol Microbiol. 2002. PMID: 11931142 Review.
-
Sure facts and open questions about the origin and evolution of photosynthetic plastids.Res Microbiol. 2001 Nov;152(9):771-80. doi: 10.1016/s0923-2508(01)01260-8. Res Microbiol. 2001. PMID: 11763237 Review.
Cited by
-
Multivesicular bodies in the enigmatic amoeboflagellate Breviata anathema and the evolution of ESCRT 0.J Cell Sci. 2011 Feb 15;124(Pt 4):613-21. doi: 10.1242/jcs.078436. Epub 2011 Jan 25. J Cell Sci. 2011. PMID: 21266469 Free PMC article.
-
Phylogeny of parasitic parabasalia and free-living relatives inferred from conventional markers vs. Rpb1, a single-copy gene.PLoS One. 2011;6(6):e20774. doi: 10.1371/journal.pone.0020774. Epub 2011 Jun 9. PLoS One. 2011. PMID: 21695260 Free PMC article.
-
Tetrahymena thermophila, a unicellular eukaryote with separate germline and somatic genomes.Res Microbiol. 2011 Jul-Aug;162(6):578-86. doi: 10.1016/j.resmic.2011.05.001. Epub 2011 May 18. Res Microbiol. 2011. PMID: 21624459 Free PMC article. Review.
-
Horizontal gene transfer of epigenetic machinery and evolution of parasitism in the malaria parasite Plasmodium falciparum and other apicomplexans.BMC Evol Biol. 2013 Feb 11;13:37. doi: 10.1186/1471-2148-13-37. BMC Evol Biol. 2013. PMID: 23398820 Free PMC article.
-
Alternative cytoskeletal landscapes: cytoskeletal novelty and evolution in basal excavate protists.Curr Opin Cell Biol. 2013 Feb;25(1):134-41. doi: 10.1016/j.ceb.2012.11.005. Epub 2013 Jan 8. Curr Opin Cell Biol. 2013. PMID: 23312067 Free PMC article. Review.
References
-
- Abascal F, Zardoya R, Posada D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics. 2005;21:2104–2105. - PubMed
-
- Adl SM, Simpson AGB, Farmer MA, Andersen RA, Anderson OR, Barta JR, Bowser SS, Brugerolle G, Fensome RA, Fredericq S, James TY, Karpov S, Kugrens P, Krug J, Lane CE, Lewis LA, Lodge J, Lynn DH, Mann DG, McCourt RM, Mendoza L, Moestrup O, Mozley-Standridge SE, Nerad TA, Shearer CA, Smirnov AV, Spiegel FW, Taylor M. The new higher level classification of eukaryotes with emphasis on the taxonomy of protists. J. Euk. Microbiol. 2005;52:399–451. - PubMed
-
- Ahmadinejad N, Dagan T, Martin W. Genome history in the symbiotic hybrid Euglena gracilis. Gene. 2007;402:35–39. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

