Regulation of alternative splicing by short non-coding nuclear RNAs
- PMID: 20657181
- PMCID: PMC3062236
- DOI: 10.4161/rna.7.4.12746
Regulation of alternative splicing by short non-coding nuclear RNAs
Abstract
Recent results from deep-sequencing and tiling array studies indicated the existence of a large number of short, metabolically stable, non-coding RNAs. Some of these short RNAs derive from known RNA classes like snoRNA or tRNAs. There are intriguing similarities between short non-coding nuclear RNAs and oligonucleotides used to change alternative splicing events, which usually target a disease-relevant RNA. We review the current knowledge of this emerging class of RNAs and discuss evidence that some of these short RNAs could function in alternative splice site selection.
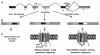
Figures

Similar articles
-
Expanding the action of duplex RNAs into the nucleus: redirecting alternative splicing.Nucleic Acids Res. 2012 Feb;40(3):1240-50. doi: 10.1093/nar/gkr780. Epub 2011 Sep 23. Nucleic Acids Res. 2012. PMID: 21948593 Free PMC article.
-
Small non-coding RNA within the endogenous spliceosome and alternative splicing regulation.Biochim Biophys Acta Gene Regul Mech. 2019 Nov-Dec;1862(11-12):194406. doi: 10.1016/j.bbagrm.2019.07.007. Epub 2019 Jul 16. Biochim Biophys Acta Gene Regul Mech. 2019. PMID: 31323432 Review.
-
Principles and correction of 5'-splice site selection.RNA Biol. 2022 Jan;19(1):943-960. doi: 10.1080/15476286.2022.2100971. RNA Biol. 2022. PMID: 35866748 Free PMC article. Review.
-
m6A modification of U6 snRNA modulates usage of two major classes of pre-mRNA 5' splice site.Elife. 2022 Nov 21;11:e78808. doi: 10.7554/eLife.78808. Elife. 2022. PMID: 36409063 Free PMC article.
-
Beyond microRNA--novel RNAs derived from small non-coding RNA and their implication in cancer.Cancer Lett. 2013 Nov 1;340(2):201-11. doi: 10.1016/j.canlet.2012.11.058. Epub 2013 Jan 29. Cancer Lett. 2013. PMID: 23376637 Review.
Cited by
-
Dyskerin Mutations Present in Dyskeratosis Congenita Patients Increase Oxidative Stress and DNA Damage Signalling in Dictyostelium Discoideum.Cells. 2019 Nov 8;8(11):1406. doi: 10.3390/cells8111406. Cells. 2019. PMID: 31717312 Free PMC article.
-
Subverting the Canon: Novel Cancer-Promoting Functions and Mechanisms for snoRNAs.Int J Mol Sci. 2024 Mar 2;25(5):2923. doi: 10.3390/ijms25052923. Int J Mol Sci. 2024. PMID: 38474168 Free PMC article. Review.
-
Role of pseudoexons and pseudointrons in human cancer.Int J Cell Biol. 2013;2013:810572. doi: 10.1155/2013/810572. Epub 2013 Sep 24. Int J Cell Biol. 2013. PMID: 24204383 Free PMC article. Review.
-
The non-coding transcriptome as a dynamic regulator of cancer metastasis.Cancer Metastasis Rev. 2014 Mar;33(1):1-16. doi: 10.1007/s10555-013-9455-3. Cancer Metastasis Rev. 2014. PMID: 24346158 Free PMC article. Review.
-
Differential evolution of signal-responsive RNA elements and upstream factors that control alternative splicing.Cell Mol Life Sci. 2014 Nov;71(22):4347-60. doi: 10.1007/s00018-014-1688-y. Epub 2014 Jul 27. Cell Mol Life Sci. 2014. PMID: 25064062 Free PMC article. Review.
References
-
- Luhrmann R, Kastner B, Bach M. Structure of spliceosomal snRNPs and their role in pre-mRNA splicing. Biochim Biophys Acta. 1990;1087:265–292. - PubMed
-
- Mattick JS, Makunin IV. Small regulatory RNAs in mammals. Hum Mol Genet. 2005;14:121–132. - PubMed
-
- Brown JW, Marshall DF, Echeverria M. Intronic noncoding RNAs and splicing. Trends Plant Sci. 2008;13:335–342. - PubMed
-
- Matera AG, Terns RM, Terns MP. Non-coding RNAs: lessons from the small nuclear and small nucleolar RNAs. Nat Rev Mol Cell Biol. 2007;8:209–220. - PubMed
-
- Steitz JA, Tycowski KT. Small RNA chaperones for ribosome biogenesis. Science. 1995;270:1626–1627. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
