A single molecular beacon probe is sufficient for the analysis of multiple nucleic acid sequences
- PMID: 20665615
- PMCID: PMC2953724
- DOI: 10.1002/cbic.201000287
A single molecular beacon probe is sufficient for the analysis of multiple nucleic acid sequences
Abstract
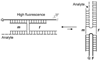
Molecular beacon (MB) probes are dual-labeled hairpin-shaped oligodeoxyribonucleotides that are extensively used for real-time detection of specific RNA/DNA analytes. In the MB probe, the loop fragment is complementary to the analyte: therefore, a unique probe is required for the analysis of each new analyte sequence. The conjugation of an oligonucleotide with two dyes and subsequent purification procedures add to the cost of MB probes, thus reducing their application in multiplex formats. Here we demonstrate how one MB probe can be used for the analysis of an arbitrary nucleic acid. The approach takes advantage of two oligonucleotide adaptor strands, each of which contains a fragment complementary to the analyte and a fragment complementary to an MB probe. The presence of the analyte leads to association of MB probe and the two DNA strands in quadripartite complex. The MB probe fluorescently reports the formation of this complex. In this design, the MB does not bind the analyte directly; therefore, the MB sequence is independent of the analyte. In this study one universal MB probe was used to genotype three human polymorphic sites. This approach promises to reduce the cost of multiplex real-time assays and improve the accuracy of single-nucleotide polymorphism genotyping.
Figures






Similar articles
-
Molecular-beacon-based tricomponent probe for SNP analysis in folded nucleic acids.Chemistry. 2011 Nov 11;17(46):13052-8. doi: 10.1002/chem.201101987. Epub 2011 Sep 28. Chemistry. 2011. PMID: 21956816 Free PMC article.
-
Operating Cooperatively (OC) sensor for highly specific recognition of nucleic acids.PLoS One. 2013;8(2):e55919. doi: 10.1371/journal.pone.0055919. Epub 2013 Feb 18. PLoS One. 2013. PMID: 23441157 Free PMC article.
-
Design and synthesis of hairpin probe for specific mis-match discrimination.Nucleic Acids Symp Ser (Oxf). 2007;(51):311-2. doi: 10.1093/nass/nrm156. Nucleic Acids Symp Ser (Oxf). 2007. PMID: 18029711
-
Single-Labeled Oligonucleotides Showing Fluorescence Changes Upon Hybridization with Target Nucleic Acids.Molecules. 2018 Jan 8;23(1):124. doi: 10.3390/molecules23010124. Molecules. 2018. PMID: 29316733 Free PMC article. Review.
-
Molecular beacons: a novel DNA probe for nucleic acid and protein studies.Chemistry. 2000 Apr 3;6(7):1107-11. doi: 10.1002/(sici)1521-3765(20000403)6:7<1107::aid-chem1107>3.3.co;2-0. Chemistry. 2000. PMID: 10785794 Review.
Cited by
-
DNA nanotechnology for nucleic acid analysis: DX motif-based sensor.Chembiochem. 2011 Nov 25;12(17):2564-7. doi: 10.1002/cbic.201100545. Epub 2011 Oct 18. Chembiochem. 2011. PMID: 22006680 Free PMC article.
-
An elegant biosensor molecular beacon probe: challenges and recent solutions.Scientifica (Cairo). 2012;2012:928783. doi: 10.6064/2012/928783. Epub 2012 Dec 13. Scientifica (Cairo). 2012. PMID: 24278758 Free PMC article. Review.
-
MicroRNA Detection Using a Double Molecular Beacon Approach: Distinguishing Between miRNA and Pre-miRNA.Theranostics. 2017 Jan 15;7(3):634-646. doi: 10.7150/thno.16840. eCollection 2017. Theranostics. 2017. PMID: 28255356 Free PMC article.
-
Molecular logic gates for DNA analysis: detection of rifampin resistance in M. tuberculosis DNA.Angew Chem Int Ed Engl. 2012 Sep 3;51(36):9075-7. doi: 10.1002/anie.201203708. Epub 2012 Aug 9. Angew Chem Int Ed Engl. 2012. PMID: 22888076 Free PMC article.
-
Molecular-beacon-based tricomponent probe for SNP analysis in folded nucleic acids.Chemistry. 2011 Nov 11;17(46):13052-8. doi: 10.1002/chem.201101987. Epub 2011 Sep 28. Chemistry. 2011. PMID: 21956816 Free PMC article.
References
-
- Morrison LE, Halder TC, Stols LM. Anal. Biochem. 1989;183:231–244. - PubMed
- Cardullo RA, Agrawal S, Flores C, Zamechnik PC, Wolf DE. Proc. Natl. Acad. Sci. USA. 1988;85:8790–8794. - PMC - PubMed
- Ebata K, Masuko M, Ohtani H, Kashiwasake-Jibu M. Photochem. Photobiol. 1995;62:836–839. - PubMed
- Santangelo P, Nitin N, Bao G. Ann. Biomed. Eng. 2006;34:39–50. - PubMed
- Silverman AP, Kool ET. Adv. Clin. Chem. Trends Biotechnol. 2005;23:225–230. - PubMed
-
- Tyagi S, Kramer FR. Nat. Biotechnol. 1996;14:303–308. - PubMed
- Marras SA, Tyagi S, Kramer FR. Clin. Chim. Acta. 2006;363:48–60. - PubMed
- Wang K, Tang Z, Yang CJ, Kim Y, Fang X, Li W, Wu Y, Medley CD, Cao Z, Li J, Colon P, Lin H, Tan W. Angew. Chem. 2009;121:870–885. - PMC - PubMed
- Angew. Chem. 2009;121:870–885.
- Angew. Chem. 2009;121:870–885.
- Li Y, Zhou X, Ye D. Biochem. Biophys. Res. Commun. 2008;373:457–461. - PubMed
- Venkatesan N, Seo YJ, Kim BH. Chem. Soc. Rev. 2008;37:648–663. - PubMed
-
- Du H, Disney MD, Miller BL, Krauss TD. J. Am. Chem. Soc. 2003;125:4012–4013. - PubMed
- Ye Y, Bloch S, Achilefu S. J. Am. Chem. Soc. 2004;126:7740–7741. - PubMed
- Liu J, Lu Y. Methods Mol. Biol. 2006;335:275–288. - PubMed
- Grossmann TN, Roglin L, Seitz O. Angew. Chem. 2007;119:5315–5318. - PubMed
- Angew. Chem. Int. Ed. 2007;46:5223–5225. - PubMed
- Lin YW, Ho HT, Huang CC, Chang HT. Nucleic Acids Res. 2008;36:e123. - PMC - PubMed
- Yamana K, Ohshita Y, Fukunaga Y, Nakamura M, Maruyama A. Bioorg. Med. Chem. 2008;16:78–83. - PubMed
- Conlon P, Yang CJ, Wu Y, Chen Y, Martinez K, Kim Y, Stevens N, Marti AA, Jockusch S, Turro NJ, Tan W. J. Am. Chem. Soc. 2008;130:336–342. - PMC - PubMed
- Matsumoto K, Shinohara Y, Bag SS, Takeuchi Y, Morii T, Saito Y, Saito I. Bioorg. Med. Chem. Lett. 2009;19:6392–6395. - PubMed
- Zhang Z, Guo L, Tang J, Guo X, Xie J. Talanta. 2009;80:985–990. - PubMed
- Nesterova IV, Erdem SS, Pakhomov S, Hammer RP, Soper SA. J. Am. Chem. Soc. 2009;131:2432–2433. - PMC - PubMed
- Häner R, Biner SM, Langenegger SM, Meng T, Malinovskii VL. Angew. Chem. 2010;122:1249–1252. - PubMed
- Angew. Chem. Int. Ed. 2010;49:1227–1230. - PubMed
- Shi C, Gu H, Ma C. Anal. Biochem. 2010;400:99–102. - PubMed
-
- Whitman DF, Dunbar SA. Recent Pat. DNA Gene Sequences. 2008;2:20–26. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

