Efficient plant gene identification based on interspecies mapping of full-length cDNAs
- PMID: 20668003
- PMCID: PMC2955710
- DOI: 10.1093/dnares/dsq017
Efficient plant gene identification based on interspecies mapping of full-length cDNAs
Abstract
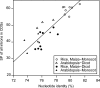
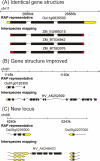
We present an annotation pipeline that accurately predicts exon-intron structures and protein-coding sequences (CDSs) on the basis of full-length cDNAs (FLcDNAs). This annotation pipeline was used to identify genes in 10 plant genomes. In particular, we show that interspecies mapping of FLcDNAs to genomes is of great value in fully utilizing FLcDNA resources whose availability is limited to several species. Because low sequence conservation at 5'- and 3'-ends of FLcDNAs between different species tends to result in truncated CDSs, we developed an improved algorithm to identify complete CDSs by the extension of both ends of truncated CDSs. Interspecies mapping of 71 801 monocot FLcDNAs to the Oryza sativa genome led to the detection of 22 142 protein-coding regions. Moreover, in comparing two mapping programs and three ab initio prediction programs, we found that our pipeline was more capable of identifying complete CDSs. As demonstrated by monocot interspecies mapping, in which nucleotide identity between FLcDNAs and the genome was ∼80%, the resultant inferred CDSs were sufficiently accurate. Finally, we applied both inter- and intraspecies mapping to 10 monocot and dicot genomes and identified genes in 210 551 loci. Interspecies mapping of FLcDNAs is expected to effectively predict genes and CDSs in newly sequenced genomes.
Figures



References
-
- Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2000;408:796–815. doi:10.1038/35048692. - DOI - PubMed
-
- International Rice Genome Sequencing Project. The map-based sequence of the rice genome. Nature. 2005;436:793–800. doi:10.1038/nature03895. - DOI - PubMed
-
- Paterson A.H., Bowers J.E., Bruggmann R., et al. The Sorghum bicolor genome and the diversification of grasses. Nature. 2009;457:551–6. doi:10.1038/nature07723. - DOI - PubMed
-
- Schnable P.S., Ware D., Fulton R.S., et al. The B73 maize genome: complexity, diversity, and dynamics. Science. 2009;326:1112–5. doi:10.1126/science.1178534. - DOI - PubMed
-
- The International Brachypodium Initiative. Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature. 2010;463:763–8. doi:10.1038/nature08747. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

