IgT, a primitive immunoglobulin class specialized in mucosal immunity
- PMID: 20676094
- PMCID: PMC3459821
- DOI: 10.1038/ni.1913
IgT, a primitive immunoglobulin class specialized in mucosal immunity
Abstract
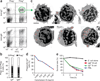
Teleost fish are the most primitive bony vertebrates that contain immunoglobulins. In contrast to mammals and birds, these species are devoid of immunoglobulin A (IgA) or a functional equivalent. This observation suggests that specialization of immunoglobulin isotypes into mucosal and systemic responses took place during tetrapod evolution. Challenging that paradigm, here we show that IgT, an immunoglobulin isotype of unknown function, acts like a mucosal antibody. We detected responses of rainbow trout IgT to an intestinal parasite only in the gut, whereas IgM responses were confined to the serum. IgT coated most intestinal bacteria. As IgT and IgA are phylogenetically distant immunoglobulins, their specialization into mucosal responses probably occurred independently by a process of convergent evolution.
Figures
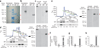



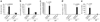


References
-
- Cooper MD, Alder MN. The evolution of adaptive immune systems. Cell. 2006;124:815–822. - PubMed
-
- Alder MN, et al. Antibody responses of variable lymphocyte receptors in the lamprey. Nat. Immunol. 2008;9:319–327. - PubMed
-
- Flajnik MF. Comparative analyses of immunoglobulin genes: surprises and portents. Nat. Rev. Immunol. 2002;2:688–698. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

