Protein kinase-inhibitor database: structural variability of and inhibitor interactions with the protein kinase P-loop
- PMID: 20681595
- PMCID: PMC3235399
- DOI: 10.1021/pr100662s
Protein kinase-inhibitor database: structural variability of and inhibitor interactions with the protein kinase P-loop
Abstract
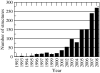
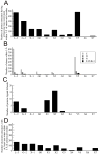
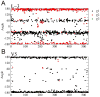
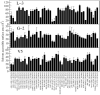
Structure-based drug design of protein-kinase inhibitors has been facilitated by availability of an enormous number of structures in the Protein Databank (PDB), systematic analyses of which can provide insight into the factors that govern ligand-protein kinase interactions and into the conformational variability of the protein kinases. In this study, a nonredundant database containing 755 unique, curated, and annotated PDB protein kinase-inhibitor complexes (each consisting of a single protein kinase chain, a ligand, and water molecules around the ligand) was created. With this dataset, analyses were performed of protein conformational variability and interactions of ligands with 11 P-loop residues. Analysis of ligand-protein interactions included ligand atom preference, ligand-protein hydrogen bonds, and the number and position of crystallographic water molecules around important P-loop residues. Analysis of variability in the conformation of the P-loop considered backbone and side-chain dihedral angles, and solvent accessible surface area (SASA). A distorted conformation of the P-loop was observed for some of the protein kinase structures. Lower SASA was observed for the hydrophobic residue in beta1 of several members of the AGC family of protein kinases. Our systematic studies were performed amino acid-by-amino acid, which is unusual for analyses of protein kinase-inhibitor complexes.
Figures
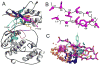
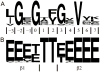


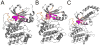
References
-
- Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S. The protein kinase complement of the human genome. Science. 2002;298(5600):1912–34. - PubMed
-
- Cherry M, Williams DH. Recent kinase and kinase inhibitor X-ray structures: mechanisms of inhibition and selectivity insights. Curr Med Chem. 2004;11(6):663–73. - PubMed
-
- Rubin GM, Yandell MD, Wortman JR, Gabor Miklos GL, Nelson CR, Hariharan IK, Fortini ME, Li PW, Apweiler R, Fleischmann W, Cherry JM, Henikoff S, Skupski MP, Misra S, Ashburner M, Birney E, Boguski MS, Brody T, Brokstein P, Celniker SE, Chervitz SA, Coates D, Cravchik A, Gabrielian A, Galle RF, Gelbart WM, George RA, Goldstein LS, Gong F, Guan P, Harris NL, Hay BA, Hoskins RA, Li J, Li Z, Hynes RO, Jones SJ, Kuehl PM, Lemaitre B, Littleton JT, Morrison DK, Mungall C, O’Farrell PH, Pickeral OK, Shue C, Vosshall LB, Zhang J, Zhao Q, Zheng XH, Lewis S. Comparative genomics of the eukaryotes. Science. 2000;287(5461):2204–15. - PMC - PubMed
-
- Huse M, Kuriyan J. The conformational plasticity of protein kinases. Cell. 2002;109(3):275–82. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

