Characterization of oligodeoxynucleotides and modifications by 193 nm photodissociation and electron photodetachment dissociation
- PMID: 20681614
- PMCID: PMC2932781
- DOI: 10.1021/ac100989q
Characterization of oligodeoxynucleotides and modifications by 193 nm photodissociation and electron photodetachment dissociation
Abstract
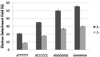
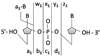
Ultraviolet photodissociation (UVPD) at 193 nm is compared to collision induced dissociation (CID) for sequencing and determination of modifications of multideprotonated 6-20-mer oligodeoxynucleotides. UVPD at 193 nm causes efficient charge reduction of the deprotonated oligodeoxynucleotides via electron detachment, in addition to extensive backbone cleavages to yield sequence ions of relatively low abundance, including w, x, y, z, a, a-B, b, c, and d ions. Although internal ions populate UVPD spectra, base loss ions from the precursor are absent. Subsequent CID of the charge-reduced oligodeoxynucleotides formed upon electron detachment, in a net process called electron photodetachment dissociation (EPD), results in abundant sequence ions in terms of w, z, a, a-B, and d products, with a marked decrease in the abundance of precursor base loss ions and internal fragments. Complete sequencing was possible for virtually all oligodeoxynucleotides studied. EPD of three modified oligodeoxynucleotides, a methylated oligodeoxynucleotide, a phosphorothioate-modified oligodeoxynucleotide, and an ethylated-oligodeoxynucleotide, resulted in specific and extensive backbone cleavages, specifically, w, z, a, a-B, and d products, which allowed the modification site(s) to be pinpointed to a more specific location than by conventional CID.
Figures






References
-
- Huber Christian, Oberacher Herbert. PCT Int. Appl. 2003:21.
-
- McLuckey ScottA, Van Berkel GaryJ, Glish GaryL. Journal of the American Society for Mass Spectrometry. 1992;3(1):60–70. - PubMed
-
- Keller KarinM, Brodbelt JenniferS. Analytical Biochemistry. 2004;326(2):200–210. - PubMed
-
- Guan Ziqiang, Kelleher NeilL, O'Connor PeterB, Aaserud DavidJ, Little DanielP, McLafferty FredW. International Journal of Mass Spectrometry and Ion Processes. 1996;157/158:357–364.
-
- Gabelica Valerie, Tabarin Thibault, Antoine Rodolphe, Rosu Frederic, Compagnon Isabelle, Broyer Michel, De Pauw Edwin, Dugourd Philippe. Analytical Chemistry. 2006;78(18):6564–6572. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

