Transient inactivation of Rb and ARF yields regenerative cells from postmitotic mammalian muscle
- PMID: 20682446
- PMCID: PMC2919350
- DOI: 10.1016/j.stem.2010.05.022
Transient inactivation of Rb and ARF yields regenerative cells from postmitotic mammalian muscle
Abstract
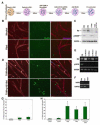
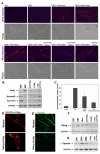
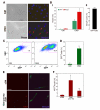
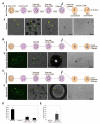
An outstanding biological question is why tissue regeneration in mammals is limited, whereas urodele amphibians and teleost fish regenerate major structures, largely by cell cycle reentry. Upon inactivation of Rb, proliferation of postmitotic urodele skeletal muscle is induced, whereas in mammalian muscle this mechanism does not exist. We postulated that a tumor suppressor present in mammals but absent in regenerative vertebrates, the Ink4a product ARF (alternative reading frame), is a regeneration suppressor. Concomitant inactivation of Arf and Rb led to mammalian muscle cell cycle reentry, loss of differentiation properties, and upregulation of cytokinetic machinery. Single postmitotic myocytes were isolated by laser micro-dissection-catapulting, and transient suppression of Arf and Rb yielded myoblast colonies that retained the ability to differentiate and fuse into myofibers upon transplantation in vivo. These results show that differentiation of mammalian cells is reversed by inactivation of Arf and Rb and support the hypothesis that Arf evolved at the expense of regeneration.
Copyright 2010 Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Newt myotubes reenter the cell cycle by phosphorylation of the retinoblastoma protein.J Cell Biol. 1997 Jan 13;136(1):155-65. doi: 10.1083/jcb.136.1.155. J Cell Biol. 1997. PMID: 9008710 Free PMC article.
-
Dual inactivation of RB and p53 pathways in RAS-induced melanomas.Mol Cell Biol. 2001 Mar;21(6):2144-53. doi: 10.1128/MCB.21.6.2144-2153.2001. Mol Cell Biol. 2001. PMID: 11238948 Free PMC article.
-
Rapid cell-cycle reentry and cell death after acute inactivation of the retinoblastoma gene product in postnatal cochlear hair cells.Proc Natl Acad Sci U S A. 2008 Jan 15;105(2):781-5. doi: 10.1073/pnas.0708061105. Epub 2008 Jan 4. Proc Natl Acad Sci U S A. 2008. PMID: 18178626 Free PMC article.
-
The Pezcoller lecture: cancer cell cycles revisited.Cancer Res. 2000 Jul 15;60(14):3689-95. Cancer Res. 2000. PMID: 10919634 Review.
-
Ink4a/Arf links senescence and aging.Exp Gerontol. 2004 Nov-Dec;39(11-12):1751-9. doi: 10.1016/j.exger.2004.06.025. Exp Gerontol. 2004. PMID: 15582292 Review.
Cited by
-
Bach1 promotes muscle regeneration through repressing Smad-mediated inhibition of myoblast differentiation.PLoS One. 2020 Aug 10;15(8):e0236781. doi: 10.1371/journal.pone.0236781. eCollection 2020. PLoS One. 2020. PMID: 32776961 Free PMC article.
-
Protein phosphatase 2A promotes the transition to G0 during terminal differentiation in Drosophila.Development. 2015 Sep 1;142(17):3033-45. doi: 10.1242/dev.120824. Epub 2015 Aug 7. Development. 2015. PMID: 26253406 Free PMC article.
-
Comparative analysis of ear-hole closure identifies epimorphic regeneration as a discrete trait in mammals.Nat Commun. 2016 Apr 25;7:11164. doi: 10.1038/ncomms11164. Nat Commun. 2016. PMID: 27109826 Free PMC article.
-
Connective tissue fibroblasts from highly regenerative mammals are refractory to ROS-induced cellular senescence.Nat Commun. 2019 Sep 27;10(1):4400. doi: 10.1038/s41467-019-12398-w. Nat Commun. 2019. PMID: 31562333 Free PMC article.
-
JARID2 and the PRC2 complex regulate the cell cycle in skeletal muscle.J Biol Chem. 2019 Dec 20;294(51):19451-19464. doi: 10.1074/jbc.RA119.010060. Epub 2019 Oct 2. J Biol Chem. 2019. PMID: 31578284 Free PMC article.
References
-
- Bartolini F, Gundersen GG. Generation of noncentrosomal microtubule arrays. Journal of cell science. 2006;119(Pt 20):4155–4163. - PubMed
-
- Bersell K, Arab S, Haring B, Kuhn B. Neuregulin1/ErbB4 signaling induces cardiomyocyte proliferation and repair of heart injury. Cell. 2009;138(2):257–270. - PubMed
-
- Blau HM, Pavlath GK, Hardeman EC, Chiu CP, Silberstein L, Webster SG, Miller SC, Webster C. Plasticity of the differentiated state. Science (New York, NY. 1985;230(4727):758–766. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R37 AG009521/AG/NIA NIH HHS/United States
- U01 HL100397/HL/NHLBI NIH HHS/United States
- T32 HD007249/HD/NICHD NIH HHS/United States
- AG009521/AG/NIA NIH HHS/United States
- T32 AI007328/AI/NIAID NIH HHS/United States
- F32 AR051678-01/AR/NIAMS NIH HHS/United States
- R01 AG020961/AG/NIA NIH HHS/United States
- R01 AG009521/AG/NIA NIH HHS/United States
- HL096113/HL/NHLBI NIH HHS/United States
- F32 AR051678/AR/NIAMS NIH HHS/United States
- AG020961/AG/NIA NIH HHS/United States
- 5T32 HD007249/HD/NICHD NIH HHS/United States
- 5T32 AI07328/AI/NIAID NIH HHS/United States
- R01 HL096113/HL/NHLBI NIH HHS/United States
- 2T32 HD007249/HD/NICHD NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

