Identification of a regulatory segment of poly(ADP-ribose) glycohydrolase
- PMID: 20684510
- PMCID: PMC2939713
- DOI: 10.1021/bi100973m
Identification of a regulatory segment of poly(ADP-ribose) glycohydrolase
Abstract
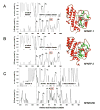
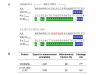
Coordinate regulation of PARP-1 and -2 and PARG is required for cellular responses to genotoxic stress. While PARP-1 and -2 are regulated by DNA breaks and covalent modifications, mechanisms of PARG regulation are poorly understood. We report here discovery of a PARG regulatory segment far removed linearly from residues involved in catalysis. Expression and analysis of human PARG segments identified a minimal catalytically active C-terminal PARG (hPARG59) containing a 16-residue N-terminal mitochondrial targeting sequence (MTS). Deletion analysis and site-directed mutagenesis revealed that the MTS, specifically hydrophobic residues L473 and L474, was required for PARG activity. This region of PARG was termed the "regulatory segment/MTS" (REG/MTS). The overall alpha-helical composition of hPARG59, determined by circular dichroism (CD), was unaffected by mutation of the REG/MTS leucine residues, suggesting that activity loss was not due to incorrect protein folding. REG/MTS was predicted to be in a loop conformation because the CD spectra of mutant Delta1-16 lacking the REG/MTS showed a higher alpha-helical content than hPARG59, indicating a secondary structure other than alpha-helix for this segment. Deletion of the REG/MTS from full-length hPARG111 also resulted in a complete loss of activity, indicating that all PARG isoforms are subject to regulation at this site. The presence of the REG/MTS raises the possibility that PARG activity is regulated by interactions of PARP-1 and -2 and other proteins at this site, raises interesting questions concerning mitochondrial PARG because MTS residues are often removed after transport, and offers a potentially novel site for drug targeting of PARG.
Figures






Similar articles
-
Two small enzyme isoforms mediate mammalian mitochondrial poly(ADP-ribose) glycohydrolase (PARG) activity.Exp Cell Res. 2007 Aug 1;313(13):2920-36. doi: 10.1016/j.yexcr.2007.03.043. Epub 2007 Apr 19. Exp Cell Res. 2007. PMID: 17509564 Free PMC article.
-
A specific isoform of poly(ADP-ribose) glycohydrolase is targeted to the mitochondrial matrix by a N-terminal mitochondrial targeting sequence.Exp Cell Res. 2009 Dec 10;315(20):3477-85. doi: 10.1016/j.yexcr.2009.04.005. Epub 2009 Apr 21. Exp Cell Res. 2009. PMID: 19389396 Free PMC article.
-
Identification of three critical acidic residues of poly(ADP-ribose) glycohydrolase involved in catalysis: determining the PARG catalytic domain.Biochem J. 2005 Jun 1;388(Pt 2):493-500. doi: 10.1042/BJ20040942. Biochem J. 2005. PMID: 15658938 Free PMC article.
-
Poly (ADP-ribose) glycohydrolase (PARG) and its therapeutic potential.Front Biosci (Landmark Ed). 2009 Jan 1;14(5):1619-26. doi: 10.2741/3329. Front Biosci (Landmark Ed). 2009. PMID: 19273151 Review.
-
New Insights into the Roles of NAD+-Poly(ADP-ribose) Metabolism and Poly(ADP-ribose) Glycohydrolase.Curr Protein Pept Sci. 2016;17(7):668-682. doi: 10.2174/1389203717666160419150014. Curr Protein Pept Sci. 2016. PMID: 27817743 Review.
Cited by
-
ADP-ribose hydrolases: biological functions and potential therapeutic targets.Expert Rev Mol Med. 2024 Oct 8;26:e21. doi: 10.1017/erm.2024.17. Expert Rev Mol Med. 2024. PMID: 39375922 Free PMC article. Review.
-
Poly(ADP-Ribose) Polymerases in Host-Pathogen Interactions, Inflammation, and Immunity.Microbiol Mol Biol Rev. 2018 Dec 19;83(1):e00038-18. doi: 10.1128/MMBR.00038-18. Print 2019 Mar. Microbiol Mol Biol Rev. 2018. PMID: 30567936 Free PMC article. Review.
-
Therapeutic Strategies and Biomarkers to Modulate PARP Activity for Targeted Cancer Therapy.Cancers (Basel). 2020 Apr 14;12(4):972. doi: 10.3390/cancers12040972. Cancers (Basel). 2020. PMID: 32295316 Free PMC article. Review.
-
Crystallographic and biochemical analysis of the mouse poly(ADP-ribose) glycohydrolase.PLoS One. 2014 Jan 21;9(1):e86010. doi: 10.1371/journal.pone.0086010. eCollection 2014. PLoS One. 2014. PMID: 24465839 Free PMC article.
-
PARG is recruited to DNA damage sites through poly(ADP-ribose)- and PCNA-dependent mechanisms.Nucleic Acids Res. 2011 Jul;39(12):5045-56. doi: 10.1093/nar/gkr099. Epub 2011 Mar 11. Nucleic Acids Res. 2011. PMID: 21398629 Free PMC article.
References
-
- Juarez-Salinas H, Sims JL, Jacobson MK. Poly(ADP-ribose) levels in carcinogen-treated cells. Nature. 1979;282:740–741. - PubMed
-
- Schreiber V, Dantzer F, Ame JC, de Murcia G. Poly(ADP-ribose): novel functions for an old molecule. Nat Rev Mol Cell Biol. 2006;7:517–528. - PubMed
-
- Heeres JT, Hergenrother PJ. Poly(ADP-ribose) makes a date with death. Curr Opin Chem Biol. 2007;11:644–653. - PubMed
-
- Kleine H, Poreba E, Lesniewicz K, Hassa PO, Hottiger MO, Litchfield DW, Shilton BH, Luscher B. Substrate-assisted catalysis by PARP10 limits its activity to mono-ADP-ribosylation. Mol Cell. 2008;32:57–69. - PubMed
-
- Hottiger MO, Hassa PO, Luscher B, Schuler H, Koch-Nolte F. Toward a unified nomenclature for mammalian ADP-ribosyltransferases. Trends Biochem Sci. 2010;35:208–219. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

