Unique cost dynamics elucidate the role of frameshifting errors in promoting translational robustness
- PMID: 20688751
- PMCID: PMC2941156
- DOI: 10.1093/gbe/evq049
Unique cost dynamics elucidate the role of frameshifting errors in promoting translational robustness
Abstract
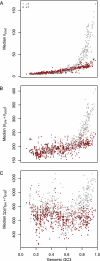
There is now considerable evidence supporting the view that codon usage is frequently under selection for translational accuracy. There are, however, multiple forms of inaccuracy (missense, premature termination, and frameshifting errors) and pinpointing a particular error process behind apparently adaptive mRNA anatomy is rarely straightforward. Understanding differences in the fitness costs associated with different types of translational error can help us devise critical tests that can implicate one error process to the exclusion of others. To this end, we present a model that captures distinct features of frameshifting cost and apply this to 641 prokaryotic genomes. We demonstrate that, although it is commonly assumed that the ribosome encounters an off-frame stop codon soon after the frameshift and costs of mis-elongation are therefore limited, genomes with high GC content typically incur much larger per-error costs. We go on to derive the prediction, unique to frameshifting errors, that differences in translational robustness between the 5' and 3' ends of genes should be less pronounced in genomes with higher GC content. This prediction we show to be correct. Surprisingly, this does not mean that GC-rich organisms necessarily carry a greater fitness burden as a consequence of accidental frameshifting. Indeed, increased per-error costs are often more than counterbalanced by lower predicted error rates owing to more diverse anticodon repertoires in GC-rich genomes. We therefore propose that selection on tRNA repertoires may operate to reduce frameshifting errors.
Figures


Similar articles
-
How translational accuracy influences reading frame maintenance.EMBO J. 1999 Mar 15;18(6):1427-34. doi: 10.1093/emboj/18.6.1427. EMBO J. 1999. PMID: 10075915 Free PMC article. Review.
-
Selection for minimization of translational frameshifting errors as a factor in the evolution of codon usage.Nucleic Acids Res. 2009 Nov;37(20):6799-810. doi: 10.1093/nar/gkp712. Epub 2009 Sep 10. Nucleic Acids Res. 2009. PMID: 19745054 Free PMC article.
-
Translational frameshifting: implications for the mechanism of translational frame maintenance.Prog Nucleic Acid Res Mol Biol. 2000;64:131-70. doi: 10.1016/s0079-6603(00)64004-7. Prog Nucleic Acid Res Mol Biol. 2000. PMID: 10697409 Review.
-
Predicting ribosomal frameshifting efficiency.Phys Biol. 2008 Mar 11;5(1):016002. doi: 10.1088/1478-3975/5/1/016002. Phys Biol. 2008. PMID: 18367782 Free PMC article.
-
Mechanism of tRNA-mediated +1 ribosomal frameshifting.Proc Natl Acad Sci U S A. 2018 Oct 30;115(44):11226-11231. doi: 10.1073/pnas.1809319115. Epub 2018 Sep 27. Proc Natl Acad Sci U S A. 2018. PMID: 30262649 Free PMC article.
Cited by
-
Adenine Enrichment at the Fourth CDS Residue in Bacterial Genes Is Consistent with Error Proofing for +1 Frameshifts.Mol Biol Evol. 2017 Dec 1;34(12):3064-3080. doi: 10.1093/molbev/msx223. Mol Biol Evol. 2017. PMID: 28961919 Free PMC article.
-
Refining the Ambush Hypothesis: Evidence That GC- and AT-Rich Bacteria Employ Different Frameshift Defence Strategies.Genome Biol Evol. 2018 Apr 1;10(4):1153-1173. doi: 10.1093/gbe/evy075. Genome Biol Evol. 2018. PMID: 29617761 Free PMC article.
-
Open questions in the study of de novo genes: what, how and why.Nat Rev Genet. 2016 Sep;17(9):567-78. doi: 10.1038/nrg.2016.78. Epub 2016 Jul 25. Nat Rev Genet. 2016. PMID: 27452112 Review.
-
In eubacteria, unlike eukaryotes, there is no evidence for selection favouring fail-safe 3' additional stop codons.PLoS Genet. 2019 Sep 17;15(9):e1008386. doi: 10.1371/journal.pgen.1008386. eCollection 2019 Sep. PLoS Genet. 2019. PMID: 31527909 Free PMC article.
-
Error prevention and mitigation as forces in the evolution of genes and genomes.Nat Rev Genet. 2011 Nov 18;12(12):875-81. doi: 10.1038/nrg3092. Nat Rev Genet. 2011. PMID: 22094950 Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

