Activation of the latent PlcR regulon in Bacillus anthracis
- PMID: 20688829
- PMCID: PMC3068694
- DOI: 10.1099/mic.0.041418-0
Activation of the latent PlcR regulon in Bacillus anthracis
Abstract
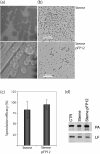
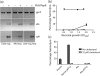
Many genes in Bacillus cereus and Bacillus thuringiensis are under the control of the transcriptional regulator PlcR and its regulatory peptide, PapR. In Bacillus anthracis, the causative agent of anthrax, PlcR is inactivated by truncation, and consequently genes having PlcR binding sites are expressed at very low levels when compared with B. cereus. We found that activation of the PlcR regulon in B. anthracis by expression of a PlcR-PapR fusion protein does not alter sporulation in strains containing the virulence plasmid pXO1 and thereby the global regulator AtxA. Using comparative 2D gel electrophoresis, we showed that activation of the PlcR regulon in B. anthracis leads to upregulation of many proteins found in the secretome of B. cereus, including phospholipases and proteases, such as the putative protease BA1995. Transcriptional analysis demonstrated expression of BA1995 to be dependent on PlcR-PapR, even though the putative PlcR recognition site of the BA1995 gene does not exactly match the PlcR consensus sequence, explaining why this protein had escaped recognition as belonging to the PlcR regulon. Additionally, while transcription of major PlcR-dependent haemolysins, sphingomyelinase and anthrolysin O is enhanced in response to PlcR activation in B. anthracis, only anthrolysin O contributes significantly to lysis of human erythrocytes. In contrast, the toxicity of bacterial culture supernatants from a PlcR-positive strain towards murine macrophages occurred independently of anthrolysin O expression in vitro and in vivo.
Figures



Similar articles
-
A spontaneous translational fusion of Bacillus cereus PlcR and PapR activates transcription of PlcR-dependent genes in Bacillus anthracis via binding with a specific palindromic sequence.Infect Immun. 2004 Oct;72(10):5814-23. doi: 10.1128/IAI.72.10.5814-5823.2004. Infect Immun. 2004. PMID: 15385482 Free PMC article.
-
The incompatibility between the PlcR- and AtxA-controlled regulons may have selected a nonsense mutation in Bacillus anthracis.Mol Microbiol. 2001 Dec;42(5):1189-98. doi: 10.1046/j.1365-2958.2001.02692.x. Mol Microbiol. 2001. PMID: 11886551
-
plcR papR-independent expression of anthrolysin O by Bacillus anthracis.J Bacteriol. 2006 Nov;188(22):7823-9. doi: 10.1128/JB.00525-06. Epub 2006 Sep 15. J Bacteriol. 2006. PMID: 16980467 Free PMC article.
-
What sets Bacillus anthracis apart from other Bacillus species?Annu Rev Microbiol. 2009;63:451-76. doi: 10.1146/annurev.micro.091208.073255. Annu Rev Microbiol. 2009. PMID: 19514852 Review.
-
AtxA, a Bacillus anthracis global virulence regulator.Res Microbiol. 2010 Nov;161(9):735-42. doi: 10.1016/j.resmic.2010.09.006. Epub 2010 Sep 21. Res Microbiol. 2010. PMID: 20863885 Review.
Cited by
-
A new quorum-sensing system (TprA/PhrA) for Streptococcus pneumoniae D39 that regulates a lantibiotic biosynthesis gene cluster.Mol Microbiol. 2015 Jul;97(2):229-43. doi: 10.1111/mmi.13029. Epub 2015 May 26. Mol Microbiol. 2015. PMID: 25869931 Free PMC article.
-
Antimicrobial Resistance Profile, Whole-Genome Sequencing and Core Genome Multilocus Sequence Typing of B. anthracis Isolates in Croatia from 2001 to 2022.Antibiotics (Basel). 2024 Jul 11;13(7):639. doi: 10.3390/antibiotics13070639. Antibiotics (Basel). 2024. PMID: 39061321 Free PMC article.
-
The Food Poisoning Toxins of Bacillus cereus.Toxins (Basel). 2021 Jan 28;13(2):98. doi: 10.3390/toxins13020098. Toxins (Basel). 2021. PMID: 33525722 Free PMC article. Review.
-
Bacterial adaptation through loss of function.PLoS Genet. 2013;9(7):e1003617. doi: 10.1371/journal.pgen.1003617. Epub 2013 Jul 11. PLoS Genet. 2013. PMID: 23874220 Free PMC article.
-
The Bacillus cereus Hbl and Nhe tripartite enterotoxin components assemble sequentially on the surface of target cells and are not interchangeable.PLoS One. 2013 Oct 18;8(10):e76955. doi: 10.1371/journal.pone.0076955. eCollection 2013. PLoS One. 2013. PMID: 24204713 Free PMC article.
References
-
- Agaisse, H., Gominet, M., Okstad, O. A., Kolstø, A. B. & Lereclus, D. (1999). PlcR is a pleiotropic regulator of extracellular virulence factor gene expression in Bacillus thuringiensis. Mol Microbiol 32, 1043–1053. - PubMed
-
- Beecher, D. J. & Wong, A. C. (2000). Cooperative, synergistic and antagonistic haemolytic interactions between haemolysin BL, phosphatidylcholine phospholipase C and sphingomyelinase from Bacillus cereus. Microbiology 146, 3033–3039. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

