Folding of matrix metalloproteinase-2 prevents endogenous generation of MHC class-I restricted epitope
- PMID: 20689590
- PMCID: PMC2912773
- DOI: 10.1371/journal.pone.0011894
Folding of matrix metalloproteinase-2 prevents endogenous generation of MHC class-I restricted epitope
Abstract
Background: We previously demonstrated that the matrix metalloproteinase-2 (MMP-2) contained an antigenic peptide recognized by a CD8 T cell clone in the HLA-A*0201 context. The presentation of this peptide on class I molecules by human melanoma cells required a cross-presentation mechanism. Surprisingly, the classical endogenous processing pathway did not process this MMP-2 epitope.
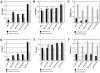
Methodology/principal findings: By PCR directed mutagenesis we showed that disruption of a single disulfide bond induced MMP-2 epitope presentation. By Pulse-Chase experiment, we demonstrated that disulfide bonds stabilized MMP-2 and impeded its degradation. Finally, using drugs, we documented that mutated MMP-2 epitope presentation used the proteasome and retrotranslocation complex.
Conclusions/significance: These data appear crucial to us since they established the existence of a new inhibitory mechanism for the generation of a T cell epitope. In spite of MMP-2 classified as a self-antigen, the fact that cross-presentation is the only way to present this MMP-2 epitope underlines the importance to target this type of antigen in immunotherapy protocols.
Conflict of interest statement
Figures





References
-
- Boel P, Wildmann C, Sensi ML, Brasseur R, Renauld JC, et al. BAGE: a new gene encoding an antigen recognized on human melanomas by cytolytic T lymphocytes. Immunity. 1995;2(2):167–175. - PubMed
-
- van der Bruggen P, Van den Eynde B. Molecular definition of tumor antigens recognized by T lymphocytes. Curr Opin Immunol. 1992;4(5):608–612. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

