RNA structural motifs that entail hydrogen bonds involving sugar-phosphate backbone atoms of RNA
- PMID: 20689681
- PMCID: PMC2915461
- DOI: 10.1039/b9nj00754g
RNA structural motifs that entail hydrogen bonds involving sugar-phosphate backbone atoms of RNA
Abstract
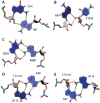
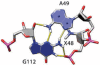
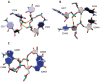
The growing number of high-resolution crystal structures of large RNA molecules provides much information for understanding the principles of structural organization of these complex molecules. Several in-depth analyses of nucleobase-centered RNA structural motifs and backbone conformations have been published based on this information, including a systematic classification of base pairs by Leontis and Westhof. However, hydrogen bonds involving sugar-phosphate backbone atoms of RNA have not been analyzed systematically until recently, although such hydrogen bonds appear to be common both in local and tertiary interactions. Here we review some backbone structural motifs discussed in the literature and analyze a set of eight high-resolution multi-domain RNA structures. The analyzed RNAs are highly structured: among 5372 nucleotides in this set, 89% are involved in at least one "long-range" RNA-RNA hydrogen bond, i.e., hydrogen bonds between atoms in the same residue or sequential residues are ignored. These long-range hydrogen bonds frequently use backbone atoms as hydrogen bond acceptors, i.e., OP1, OP2, O2', O3', O4', or O5', or as a donor (2'OH). A surprisingly large number of such hydrogen bonds are found, considering that neither single-stranded nor double-stranded regions will contain such hydrogen bonds unless additional interactions with other residues exist. Among 8327 long-range hydrogen bonds found in this set of structures, 2811, or about one-third, are hydrogen bonds entailing RNA backbone atoms; they involve 39% of all nucleotides in the structures. The majority of them (2111) are hydrogen bonds entailing ribose hydroxyl groups, which can be used either as a donor or an acceptor; they constitute 25% of all hydrogen bonds and involve 31% of all nucleotides. The phosphate oxygens OP1 or OP2 are used as hydrogen bond acceptors in 12% of all nucleotides, and the ribose ring oxygen O4' and phosphodiester oxygens O3' and O5' are used in 4%, 4%, and 1% of all nucleotides, respectively. Distributions of geometric parameters and some examples of such hydrogen bonds are presented in this report. A novel motif involving backbone hydrogen bonds, the ribose-phosphate zipper, is also identified.
Figures

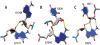

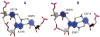

Similar articles
-
A molecular dynamics simulation of the flavin mononucleotide-RNA aptamer complex.Biopolymers. 1999 Sep;50(3):287-302. doi: 10.1002/(SICI)1097-0282(199909)50:3<287::AID-BIP5>3.0.CO;2-G. Biopolymers. 1999. PMID: 10397790
-
Crystal structure of RNase T1 complexed with the product nucleotide 3'-GMP. Structural evidence for direct interaction of histidine 40 and glutamic acid 58 with the 2'-hydroxyl group of the ribose.J Biol Chem. 1994 Jul 1;269(26):17531-6. doi: 10.2210/pdb1rls/pdb. J Biol Chem. 1994. PMID: 7912696
-
The significance of the 2' OH group and the influence of cations on the secondary structure of the RNA backbone.Biophys Struct Mech. 1975 May 30;1(3):203-19. doi: 10.1007/BF00535757. Biophys Struct Mech. 1975. PMID: 1234026
-
A structural role for arginine in proteins: multiple hydrogen bonds to backbone carbonyl oxygens.Protein Sci. 1994 Apr;3(4):541-8. doi: 10.1002/pro.5560030402. Protein Sci. 1994. PMID: 8003972 Free PMC article. Review.
-
Gabapentin: a stereochemically constrained gamma amino acid residue in hybrid peptide design.Acc Chem Res. 2009 Oct 20;42(10):1628-39. doi: 10.1021/ar9001153. Acc Chem Res. 2009. PMID: 19572698 Review.
Cited by
-
NMR structures and magnetic force spectroscopy studies of small molecules binding to models of an RNA CAG repeat expansion.bioRxiv [Preprint]. 2024 Aug 21:2024.08.20.608150. doi: 10.1101/2024.08.20.608150. bioRxiv. 2024. PMID: 39229124 Free PMC article. Preprint.
-
Recognition of RNA duplexes by chemically modified triplex-forming oligonucleotides.Nucleic Acids Res. 2013 Jul;41(13):6664-73. doi: 10.1093/nar/gkt352. Epub 2013 May 8. Nucleic Acids Res. 2013. PMID: 23658228 Free PMC article.
-
Carbon nanospikes have improved sensitivity and antifouling properties for adenosine, hydrogen peroxide, and histamine.Anal Bioanal Chem. 2023 Oct;415(24):6039-6050. doi: 10.1007/s00216-023-04875-5. Epub 2023 Jul 28. Anal Bioanal Chem. 2023. PMID: 37505236 Free PMC article.
-
RNA: The Unsuspected Conductor in the Orchestra of Macromolecular Crowding.Chem Rev. 2024 Apr 24;124(8):4734-4777. doi: 10.1021/acs.chemrev.3c00575. Epub 2024 Apr 5. Chem Rev. 2024. PMID: 38579177 Free PMC article. Review.
-
Crystal structure of a cap-independent translation enhancer RNA.Nucleic Acids Res. 2023 Sep 8;51(16):8891-8907. doi: 10.1093/nar/gkad649. Nucleic Acids Res. 2023. PMID: 37548413 Free PMC article.
References
-
- Holbrook SR. Annu. Rev. Biophys. Biomol. Struct. 2008;37:445–464. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
