Global analyses of small interfering RNAs derived from Bamboo mosaic virus and its associated satellite RNAs in different plants
- PMID: 20689857
- PMCID: PMC2914070
- DOI: 10.1371/journal.pone.0011928
Global analyses of small interfering RNAs derived from Bamboo mosaic virus and its associated satellite RNAs in different plants
Abstract
Background: Satellite RNAs (satRNAs), virus parasites, are exclusively associated with plant virus infection and have attracted much interest over the last 3 decades. Upon virus infection, virus-specific small interfering RNAs (vsiRNAs) are produced by dicer-like (DCL) endoribonucleases for anti-viral defense. The composition of vsiRNAs has been studied extensively; however, studies of satRNA-derived siRNAs (satsiRNAs) or siRNA profiles after satRNA co-infection are limited. Here, we report on the small RNA profiles associated with infection with Bamboo mosaic virus (BaMV) and its two satellite RNAs (satBaMVs) in Nicotiana benthamiana and Arabidopsis thaliana.
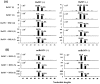
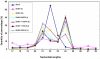
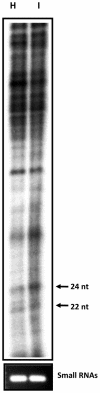
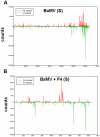
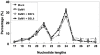
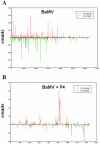
Methodology/principal findings: Leaves of N. benthamiana or A. thaliana inoculated with water, BaMV alone or co-inoculated with interfering or noninterfering satBaMV were collected for RNA extraction, then large-scale Solexa sequencing. Up to about 20% of total siRNAs as BaMV-specific siRNAs were accumulated in highly susceptible N. benthamiana leaves inoculated with BaMV alone or co-inoculated with noninterfering satBaMV; however, only about 0.1% of vsiRNAs were produced in plants co-infected with interfering satBaMV. The abundant region of siRNA distribution along BaMV and satBaMV genomes differed by host but not by co-infection with satBaMV. Most of the BaMV and satBaMV siRNAs were 21 or 22 nt, of both (+) and (-) polarities; however, a higher proportion of 22-nt BaMV and satBaMV siRNAs were generated in N. benthamiana than in A. thaliana. Furthermore, the proportion of non-viral 24-nt siRNAs was greatly increased in N. benthamiana after virus infection.
Conclusions/significance: The overall composition of vsiRNAs and satsiRNAs in the infected plants reflect the combined action of virus, satRNA and different DCLs in host plants. Our findings suggest that the structure and/or sequence demands of various DCLs in different hosts may result in differential susceptibility to the same virus. DCL2 producing 24-nt siRNAs under biotic stresses may play a vital role in the antiviral mechanism in N. benthamiana.
Conflict of interest statement
Figures



Similar articles
-
Transgenic resistance to Bamboo mosaic virus by expression of interfering satellite RNA.Mol Plant Pathol. 2013 Sep;14(7):693-707. doi: 10.1111/mpp.12040. Epub 2013 May 16. Mol Plant Pathol. 2013. PMID: 23675895 Free PMC article.
-
Characterisation of siRNAs derived from new isolates of bamboo mosaic virus and their associated satellites in infected ma bamboo (Dendrocalamus latiflorus).Arch Virol. 2017 Feb;162(2):505-510. doi: 10.1007/s00705-016-3092-4. Epub 2016 Oct 14. Arch Virol. 2017. PMID: 27743256
-
Generation of subgenomic RNA directed by a satellite RNA associated with bamboo mosaic potexvirus: analyses of potexvirus subgenomic RNA promoter.J Virol. 2000 Nov;74(22):10341-8. doi: 10.1128/jvi.74.22.10341-10348.2000. J Virol. 2000. PMID: 11044078 Free PMC article.
-
Interfering Satellite RNAs of Bamboo mosaic virus.Front Microbiol. 2017 May 4;8:787. doi: 10.3389/fmicb.2017.00787. eCollection 2017. Front Microbiol. 2017. PMID: 28522996 Free PMC article. Review.
-
Viral elements and host cellular proteins in intercellular movement of Bamboo mosaic virus.Curr Opin Virol. 2015 Jun;12:99-108. doi: 10.1016/j.coviro.2015.04.005. Epub 2015 May 13. Curr Opin Virol. 2015. PMID: 25951346 Review.
Cited by
-
The Tomato Spotted Wilt Virus Genome Is Processed Differentially in its Plant Host Arachis hypogaea and its Thrips Vector Frankliniella fusca.Front Plant Sci. 2016 Sep 7;7:1349. doi: 10.3389/fpls.2016.01349. eCollection 2016. Front Plant Sci. 2016. PMID: 27656190 Free PMC article.
-
Genome-wide analysis of small RNAs from Odontoglossum ringspot virus and Cymbidium mosaic virus synergistically infecting Phalaenopsis.Mol Plant Pathol. 2020 Feb;21(2):188-205. doi: 10.1111/mpp.12888. Epub 2019 Nov 14. Mol Plant Pathol. 2020. PMID: 31724809 Free PMC article.
-
Dual resistance of transgenic plants against Cymbidium mosaic virus and Odontoglossum ringspot virus.Sci Rep. 2019 Jul 15;9(1):10230. doi: 10.1038/s41598-019-46695-7. Sci Rep. 2019. PMID: 31308424 Free PMC article.
-
MiR398-regulated antioxidants contribute to Bamboo mosaic virus accumulation and symptom manifestation.Plant Physiol. 2022 Jan 20;188(1):593-607. doi: 10.1093/plphys/kiab451. Plant Physiol. 2022. PMID: 34695209 Free PMC article.
-
Differential expression of tomato spotted wilt virus-derived viral small RNAs in infected commercial and experimental host plants.PLoS One. 2013 Oct 15;8(10):e76276. doi: 10.1371/journal.pone.0076276. eCollection 2013. PLoS One. 2013. PMID: 24143182 Free PMC article.
References
-
- Voinnet O. RNA silencing as a plant immune system against viruses. Trends Genet. 2001;17:449–459. - PubMed
-
- Ding SW, Li H, Lu R, Li F, Li WX. RNA silencing: a conserved antiviral immunity of plants and animals. Virus Res. 2004;102:109–115. - PubMed
-
- Waterhouse PM, Wang MB, Lough T. Gene silencing as an adaptive defence against viruses. Nature. 2001;411:834–842. - PubMed
-
- Vance V, Vaucheret H. RNA silencing in plants–defense and counterdefense. Science. 2001;292:2277–2280. - PubMed
-
- Baulcombe D. RNA silencing in plants. Nature. 2004;431:356–363. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

