Flow cytometry analysis of transcription factors in T lymphocytes
- PMID: 20694680
- PMCID: PMC3788577
- DOI: 10.1007/978-1-60761-738-9_23
Flow cytometry analysis of transcription factors in T lymphocytes
Abstract
Detection of transcription factors in immune cell populations, particularly in subpopulations that are represented at low frequencies in lymphoid and nonlymphoid organs, presents a particular challenge when using traditional methods such as western blot analysis. Therefore, development of flow cytometry-based methods which allow identification of transcription factors in specific immune cell populations is of main interest. Here we developed and optimized a methodology for rapid and convenient detection of the transcription factor BCL11B in T lymphocyte subpopulations using flow cytometry. The optimal protocol employs saponin and Tween 20 both during the fixation and permeabilization steps, and we demonstrate that it is efficient for three anti-BCL11B antibodies covering distinctive BCL11B epitopes. In addition, we prove that the method preserves the staining of surface markers.
Figures



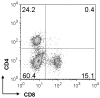
References
-
- Fontenot JD, Rasmussen JP, Williams LM, Dooley JL, Farr AG, Rudensky AY. Regulatory T cell lineage specification by the forkhead transcription factor foxp3. Immunity. 2005;22(3):329–41. - PubMed
-
- Ivanov, McKenzie BS, Zhou L, et al. The orphan nuclear receptor RORgammat directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell. 2006;126(6):1121–33. - PubMed
-
- Wakabayashi Y, Watanabe H, Inoue J, et al. Bcl11b is required for differentiation and survival of alphabeta T lymphocytes. Nat Immunol. 2003;4(6):533–9. - PubMed
-
- Cismasiu VB, Adamo K, Gecewicz J, Duque J, Lin Q, Avram D. BCL11B functionally associates with the NuRD complex in T lymphocytes to repress targeted promoter. Oncogene. 2005;24(45):6753–64. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

