Sequence-specific cleavage of RNA by Type II restriction enzymes
- PMID: 20702422
- PMCID: PMC3001074
- DOI: 10.1093/nar/gkq702
Sequence-specific cleavage of RNA by Type II restriction enzymes
Abstract
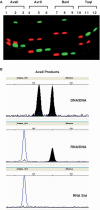
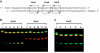
The ability of 223 Type II restriction endonucleases to hydrolyze RNA-DNA heteroduplex oligonucleotide substrates was assessed. Despite the significant topological and sequence asymmetry introduced when one strand of a DNA duplex is substituted by RNA we find that six restriction enzymes (AvaII, AvrII, BanI, HaeIII, HinfI and TaqI), exclusively of the Type IIP class that recognize palindromic or interrupted-palindromic DNA sequences, catalyze robust and specific cleavage of both RNA and DNA strands of such a substrate. Time-course analyses indicate that some endonucleases hydrolyze phosphodiester bonds in both strands simultaneously whereas others appear to catalyze sequential reactions in which either the DNA or RNA product accumulates more rapidly. Such strand-specific variation in cleavage susceptibility is both significant (up to orders of magnitude difference) and somewhat sequence dependent, notably in relation to the presence or absence of uracil residues in the RNA strand. Hybridization to DNA oligonucleotides that contain endonuclease recognition sites can be used to achieve targeted hydrolysis of extended RNA substrates produced by in vitro transcription. The ability to 'restrict' an RNA-DNA hybrid, albeit with a limited number of restriction endonucleases, provides a method whereby individual RNA molecules can be targeted for site-specific cleavage in vitro.
Figures




Similar articles
-
Target site cleavage by the monomeric restriction enzyme BcnI requires translocation to a random DNA sequence and a switch in enzyme orientation.Nucleic Acids Res. 2011 Nov 1;39(20):8844-56. doi: 10.1093/nar/gkr588. Epub 2011 Jul 19. Nucleic Acids Res. 2011. PMID: 21771860 Free PMC article.
-
Oligonucleotide cleavage by restriction endonucleases MvaI and EcoRII: a comprehensive study on the influence of structural parameters on the enzyme-substrate interaction.Biochim Biophys Acta. 1991 Mar 26;1088(3):395-400. doi: 10.1016/0167-4781(91)90131-5. Biochim Biophys Acta. 1991. PMID: 2015301
-
Restriction endonuclease AgeI is a monomer which dimerizes to cleave DNA.Nucleic Acids Res. 2017 Apr 7;45(6):3547-3558. doi: 10.1093/nar/gkw1310. Nucleic Acids Res. 2017. PMID: 28039325 Free PMC article.
-
Recognition and cleavage of DNA by type-II restriction endonucleases.Eur J Biochem. 1997 May 15;246(1):1-22. doi: 10.1111/j.1432-1033.1997.t01-6-00001.x. Eur J Biochem. 1997. PMID: 9210460 Review.
-
Natural and engineered nicking endonucleases--from cleavage mechanism to engineering of strand-specificity.Nucleic Acids Res. 2011 Jan;39(1):1-18. doi: 10.1093/nar/gkq742. Epub 2010 Aug 30. Nucleic Acids Res. 2011. PMID: 20805246 Free PMC article. Review.
Cited by
-
Opportunities for Expanding Encoded Chemical Diversification and Improving Hit Enrichment in mRNA-Displayed Peptide Libraries.Chembiochem. 2022 Jun 20;23(12):e202100685. doi: 10.1002/cbic.202100685. Epub 2022 Feb 18. Chembiochem. 2022. PMID: 35100479 Free PMC article. Review.
-
Highlights of the DNA cutters: a short history of the restriction enzymes.Nucleic Acids Res. 2014 Jan;42(1):3-19. doi: 10.1093/nar/gkt990. Epub 2013 Oct 18. Nucleic Acids Res. 2014. PMID: 24141096 Free PMC article.
-
Relaxed specificity of prokaryotic DNA methyltransferases results in DNA site-specific modification of RNA/DNA heteroduplexes.J Appl Genet. 2015 Nov;56(4):539-546. doi: 10.1007/s13353-015-0279-4. Epub 2015 Mar 19. J Appl Genet. 2015. PMID: 25787880
-
Types and Applications of Nicking Enzyme-Combined Isothermal Amplification.Int J Mol Sci. 2022 Apr 21;23(9):4620. doi: 10.3390/ijms23094620. Int J Mol Sci. 2022. PMID: 35563012 Free PMC article. Review.
-
Digital imprinting of RNA recognition and processing on a self-assembled nucleic acid matrix.Sci Rep. 2013;3:2550. doi: 10.1038/srep02550. Sci Rep. 2013. PMID: 23989631 Free PMC article.
References
-
- Smith HO, Wilcox KW. A restriction enzyme from Hemophilus influenzae. I. Purification and general properties. J. Mol. Biol. 1970;51:379–391. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

