Structural basis for the bacterial transcription-repair coupling factor/RNA polymerase interaction
- PMID: 20702425
- PMCID: PMC3001067
- DOI: 10.1093/nar/gkq692
Structural basis for the bacterial transcription-repair coupling factor/RNA polymerase interaction
Abstract
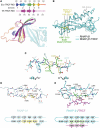
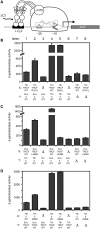
The transcription-repair coupling factor (TRCF, the product of the mfd gene) is a widely conserved bacterial protein that mediates transcription-coupled DNA repair. TRCF uses its ATP-dependent DNA translocase activity to remove transcription complexes stalled at sites of DNA damage, and stimulates repair by recruiting components of the nucleotide excision repair pathway to the site. A protein/protein interaction between TRCF and the β-subunit of RNA polymerase (RNAP) is essential for TRCF function. CarD (also called CdnL), an essential regulator of rRNA transcription in Mycobacterium tuberculosis, shares a homologous RNAP interacting domain with TRCF and also interacts with the RNAP β-subunit. We determined the 2.9-Å resolution X-ray crystal structure of the RNAP interacting domain of TRCF complexed with the RNAP-β1 domain, which harbors the TRCF interaction determinants. The structure reveals details of the TRCF/RNAP protein/protein interface, providing a basis for the design and interpretation of experiments probing TRCF, and by homology CarD, function and interactions with the RNAP.
Figures
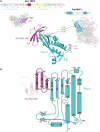

Similar articles
-
Structure-function dissection of Myxococcus xanthus CarD N-terminal domain, a defining member of the CarD_CdnL_TRCF family of RNA polymerase interacting proteins.PLoS One. 2015 Mar 26;10(3):e0121322. doi: 10.1371/journal.pone.0121322. eCollection 2015. PLoS One. 2015. PMID: 25811865 Free PMC article.
-
Structure of the Mtb CarD/RNAP β-lobes complex reveals the molecular basis of interaction and presents a distinct DNA-binding domain for Mtb CarD.Structure. 2013 Oct 8;21(10):1859-69. doi: 10.1016/j.str.2013.08.014. Epub 2013 Sep 19. Structure. 2013. PMID: 24055315 Free PMC article.
-
Structural basis for bacterial transcription-coupled DNA repair.Cell. 2006 Feb 10;124(3):507-20. doi: 10.1016/j.cell.2005.11.045. Cell. 2006. PMID: 16469698
-
The bacterial transcription repair coupling factor.Curr Opin Struct Biol. 2007 Feb;17(1):96-102. doi: 10.1016/j.sbi.2007.01.005. Epub 2007 Jan 18. Curr Opin Struct Biol. 2007. PMID: 17239578 Free PMC article. Review.
-
Mfd Protein and Transcription-Repair Coupling in Escherichia coli.Photochem Photobiol. 2017 Jan;93(1):280-295. doi: 10.1111/php.12675. Epub 2017 Jan 18. Photochem Photobiol. 2017. PMID: 27864884 Free PMC article. Review.
Cited by
-
Cotranscriptional R-loop formation by Mfd involves topological partitioning of DNA.Proc Natl Acad Sci U S A. 2021 Apr 13;118(15):e2019630118. doi: 10.1073/pnas.2019630118. Proc Natl Acad Sci U S A. 2021. PMID: 33827922 Free PMC article.
-
Helicases as transcription termination factors: Different solutions for a common problem.Transcription. 2018;9(3):152-158. doi: 10.1080/21541264.2017.1361503. Epub 2017 Oct 4. Transcription. 2018. PMID: 28886303 Free PMC article.
-
Structure of the carboxy-terminal domain of Mycobacterium tuberculosis CarD protein: an essential rRNA transcriptional regulator.Acta Crystallogr F Struct Biol Commun. 2014 Feb;70(Pt 2):160-5. doi: 10.1107/S2053230X13034407. Epub 2014 Jan 21. Acta Crystallogr F Struct Biol Commun. 2014. PMID: 24637748 Free PMC article.
-
Mechanistic insights into transcription coupled DNA repair.DNA Repair (Amst). 2017 Aug;56:42-50. doi: 10.1016/j.dnarep.2017.06.006. Epub 2017 Jun 9. DNA Repair (Amst). 2017. PMID: 28629777 Free PMC article. Review.
-
The conserved C-terminus of the PcrA/UvrD helicase interacts directly with RNA polymerase.PLoS One. 2013 Oct 16;8(10):e78141. doi: 10.1371/journal.pone.0078141. eCollection 2013. PLoS One. 2013. PMID: 24147116 Free PMC article.
References
-
- Bohr VA, Smith CA, Okumoto DS, Hanawalt PC. DNA repair in an active gene: removal of pyrimidine dimers from the DHFR gene of CHO cells is much more efficient than in the genome overall. Cell. 1985;40:359–369. - PubMed
-
- Mellon I, Hanawalt PC. Induction of the Escherichia coli lactose operon selectively increases repair of its transcribed DNA strand. Nature. 1989;342:95–98. - PubMed
-
- Selby CP, Sancar A. Transcription preferentially inhibits nucleotide excision repair of the template DNA strand in vitro. J. Biol. Chem. 1990;265:21330–21336. - PubMed
-
- Witkin EM. Time, temperature, and protein synthesis: a study of ultraviolet-induced mutation in bacteria. Cold Spring Harbor Symp. Quant. Biol. 1956;21:123–140. - PubMed
-
- Witkin EM. Radiation-induced mutations and their repair. Science. 1966;152:1345–1353. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions

