UV-B induced transcript accumulation of DAHP synthase in suspension-cultured Catharanthus roseus cells
- PMID: 20704760
- PMCID: PMC2930624
- DOI: 10.1186/1750-2187-5-13
UV-B induced transcript accumulation of DAHP synthase in suspension-cultured Catharanthus roseus cells
Abstract
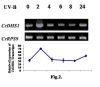
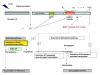
The enzyme 3-deoxy-D-arabino-heptulosonate-7-phosphate (DAHP) synthase (EC 4.1.2.15) catalyzes the first committed step in the shikimate pathway of tryptophan synthesis, an important precursor for the production of terpenoid indole alkaloids (TIAs). A full-length cDNA encoding nuclear coded chloroplast-specific DAHP synthase transcript was isolated from a Catharanthus roseus cDNA library. This had high sequence similarity with other members of plant DAHP synthase family. This transcript accumulated in suspension cultured C. roseus cells on ultraviolet (UV-B) irradiation. Pretreatment of C.roseus cells with variety of agents such as suramin, N-acetyl cysteine, and inhibitors of calcium fluxes and protein kinases and MAP kinase prevented this effect of UV-B irriadiation. These data further show that the essential components of the signaling pathway involved in accumulation DAHP synthase transcript in C. roseus cells include suramin-sensitive cell surface receptor, staurosporine-sensitive protein kinase and MAP kinase.
Figures


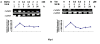


Similar articles
-
UV-B-induced signaling events leading to enhanced-production of catharanthine in Catharanthus roseus cell suspension cultures.BMC Plant Biol. 2007 Nov 7;7:61. doi: 10.1186/1471-2229-7-61. BMC Plant Biol. 2007. PMID: 17988378 Free PMC article.
-
Impaired Wound Induction of 3-Deoxy-D-arabino-heptulosonate-7-phosphate (DAHP) Synthase and Altered Stem Development in Transgenic Potato Plants Expressing a DAHP Synthase Antisense Construct.Plant Physiol. 1995 Aug;108(4):1413-1421. doi: 10.1104/pp.108.4.1413. Plant Physiol. 1995. PMID: 12228551 Free PMC article.
-
Translocation of 3-deoxy-D-arabino-heptulosonate 7-phosphate synthase precursor into isolated chloroplasts.Planta. 2002 Nov;216(1):180-6. doi: 10.1007/s00425-002-0891-5. Epub 2002 Nov 12. Planta. 2002. PMID: 12430029
-
3-Deoxy-D-arabino-heptulosonate 7-phosphate synthase as the gatekeeper of plant aromatic natural product biosynthesis.Curr Opin Plant Biol. 2022 Jun;67:102219. doi: 10.1016/j.pbi.2022.102219. Epub 2022 May 10. Curr Opin Plant Biol. 2022. PMID: 35550985 Review.
-
Production and metabolic engineering of terpenoid indole alkaloids in cell cultures of the medicinal plant Catharanthus roseus (L.) G. Don (Madagascar periwinkle).Biotechnol Appl Biochem. 2009 Apr;52(Pt 4):313-23. doi: 10.1042/BA20080239. Biotechnol Appl Biochem. 2009. PMID: 19281450 Review.
References
LinkOut - more resources
Full Text Sources

