Resistance to integrase inhibitors
- PMID: 20706558
- PMCID: PMC2920056
- DOI: 10.3390/v2071347
Resistance to integrase inhibitors
Abstract
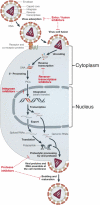
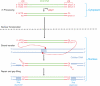
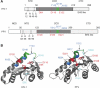
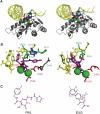
Integrase (IN) is a clinically validated target for the treatment of human immunodeficiency virus infections and raltegravir exhibits remarkable clinical activity. The next most advanced IN inhibitor is elvitegravir. However, mutant viruses lead to treatment failure and mutations within the IN coding sequence appear to confer cross-resistance. The characterization of those mutations is critical for the development of second generation IN inhibitors to overcome resistance. This review focuses on IN resistance based on structural and biochemical data, and on the role of the IN flexible loop i.e., between residues G140-G149 in drug action and resistance.
Keywords: AIDS; Elvitegravir; GSK-1265744; GSK-1349572; HIV-1 integrase; Raltegravir; interfacial inhibitors; resistance.
Figures

References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
