Transcriptomic analysis of Escherichia coli O157:H7 and K-12 cultures exposed to inorganic and organic acids in stationary phase reveals acidulant- and strain-specific acid tolerance responses
- PMID: 20709847
- PMCID: PMC2950450
- DOI: 10.1128/AEM.02392-09
Transcriptomic analysis of Escherichia coli O157:H7 and K-12 cultures exposed to inorganic and organic acids in stationary phase reveals acidulant- and strain-specific acid tolerance responses
Abstract
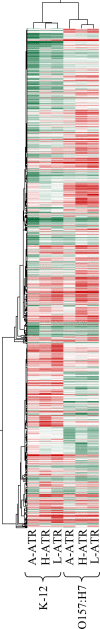
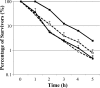
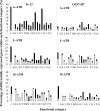
The food-borne pathogen Escherichia coli O157:H7 is commonly exposed to organic acid in processed and preserved foods, allowing adaptation and the development of tolerance to pH levels otherwise lethal. Since little is known about the molecular basis of adaptation of E. coli to organic acids, we studied K-12 MG1655 and O157:H7 Sakai during exposure to acetic, lactic, and hydrochloric acid at pH 5.5. This is the first analysis of the pH-dependent transcriptomic response of stationary-phase E. coli. Thirty-four genes and three intergenic regions were upregulated by both strains during exposure to all acids. This universal acid response included genes involved in oxidative, envelope, and cold stress resistance and iron and manganese uptake, as well as 10 genes of unknown function. Acidulant- and strain-specific responses were also revealed. The acidulant-specific response reflects differences in the modes of microbial inactivation, even between weak organic acids. The two strains exhibited similar responses to lactic and hydrochloric acid, while the response to acetic acid was distinct. Acidulant-dependent differences between the strains involved induction of genes involved in the heat shock response, osmoregulation, inorganic ion and nucleotide transport and metabolism, translation, and energy production. E. coli O157:H7-specific acid-inducible genes were identified, suggesting that the enterohemorrhagic E. coli strain possesses additional molecular mechanisms contributing to acid resistance that are absent in K-12. While E. coli K-12 was most resistant to lactic and hydrochloric acid, O157:H7 may have a greater ability to survive in more complex acidic environments, such as those encountered in the host and during food processing.
Figures

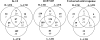
Similar articles
-
Evolutionary silence of the acid chaperone protein HdeB in enterohemorrhagic Escherichia coli O157:H7.Appl Environ Microbiol. 2012 Feb;78(4):1004-14. doi: 10.1128/AEM.07033-11. Epub 2011 Dec 16. Appl Environ Microbiol. 2012. PMID: 22179243 Free PMC article.
-
Stress Resistance Development and Genome-Wide Transcriptional Response of Escherichia coli O157:H7 Adapted to Sublethal Thymol, Carvacrol, and trans-Cinnamaldehyde.Appl Environ Microbiol. 2018 Oct 30;84(22):e01616-18. doi: 10.1128/AEM.01616-18. Print 2018 Nov 15. Appl Environ Microbiol. 2018. PMID: 30217837 Free PMC article.
-
Identifying the mechanism of Escherichia coli O157:H7 survival by the addition of salt in the treatment with organic acids.J Appl Microbiol. 2018 Jan;124(1):241-253. doi: 10.1111/jam.13613. Epub 2017 Nov 27. J Appl Microbiol. 2018. PMID: 29044827
-
Evaluation of the pH-dependent, stationary-phase acid tolerance in Listeria monocytogenes and Salmonella Typhimurium DT104 induced by culturing in media with 1% glucose: a comparative study with Escherichia coli O157:H7.J Appl Microbiol. 2003;95(3):563-75. doi: 10.1046/j.1365-2672.2003.02013.x. J Appl Microbiol. 2003. PMID: 12911705
-
Molecular Profiling: Catecholamine Modulation of Gene Expression in Escherichia coli O157:H7 and Salmonella enterica Serovar Typhimurium.Adv Exp Med Biol. 2016;874:167-82. doi: 10.1007/978-3-319-20215-0_7. Adv Exp Med Biol. 2016. PMID: 26589218 Review.
Cited by
-
13C-Metabolic Flux Analysis: An Accurate Approach to Demystify Microbial Metabolism for Biochemical Production.Bioengineering (Basel). 2015 Dec 25;3(1):3. doi: 10.3390/bioengineering3010003. Bioengineering (Basel). 2015. PMID: 28952565 Free PMC article. Review.
-
Investigation of Escherichia coli O157:H7 Survival and Interaction with Meal Components during Gastrointestinal Digestion.Foods. 2021 Oct 12;10(10):2415. doi: 10.3390/foods10102415. Foods. 2021. PMID: 34681467 Free PMC article.
-
Expanded roles of lactate-sensing LldR in transcription regulation of the Escherichia coli K-12 genome: lactate utilisation and acid resistance.Microb Genom. 2023 May;9(5):mgen001015. doi: 10.1099/mgen.0.001015. Microb Genom. 2023. PMID: 37219924 Free PMC article.
-
Evolutionary silence of the acid chaperone protein HdeB in enterohemorrhagic Escherichia coli O157:H7.Appl Environ Microbiol. 2012 Feb;78(4):1004-14. doi: 10.1128/AEM.07033-11. Epub 2011 Dec 16. Appl Environ Microbiol. 2012. PMID: 22179243 Free PMC article.
-
Stress Resistance Development and Genome-Wide Transcriptional Response of Escherichia coli O157:H7 Adapted to Sublethal Thymol, Carvacrol, and trans-Cinnamaldehyde.Appl Environ Microbiol. 2018 Oct 30;84(22):e01616-18. doi: 10.1128/AEM.01616-18. Print 2018 Nov 15. Appl Environ Microbiol. 2018. PMID: 30217837 Free PMC article.
References
-
- Antal, M., V. Bordeau, V. Douchin, and B. Felden. 2005. A small bacterial RNA regulates a putative ABC transporter. J. Biol. Chem. 280:7901-7908. - PubMed
-
- Antón, J., A. Oren, S. Benlloch, F. Rodríguez-Valera, R. Amann, and R. Rosselló-Mora. 2002. Salinibacter ruber gen. nov., sp. nov., a novel, extremely halophilic member of the Bacteria from saltern crystallizer ponds. Int. J. Syst. Evol. Microbiol. 52:485-491. - PubMed
-
- Argaman, L., R. Hershberg, J. Vogel, G. Bejerano, E. G. H. Wagner, H. Margalit, and S. Altuvia. 2001. Novel small RNA-encoding genes in the intergenic regions of Escherichia coli. Curr. Biol. 11:941-950. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

