Viral entry and escape from antibody-mediated neutralization influence hepatitis C virus reinfection in liver transplantation
- PMID: 20713596
- PMCID: PMC2931157
- DOI: 10.1084/jem.20090766
Viral entry and escape from antibody-mediated neutralization influence hepatitis C virus reinfection in liver transplantation
Abstract
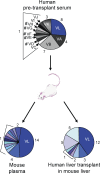
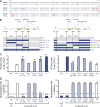
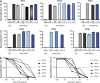
End-stage liver disease caused by chronic hepatitis C virus (HCV) infection is a leading cause for liver transplantation (LT). Due to viral evasion from host immune responses and the absence of preventive antiviral strategies, reinfection of the graft is universal. The mechanisms by which the virus evades host immunity to reinfect the liver graft are unknown. In a longitudinal analysis of six HCV-infected patients undergoing LT, we demonstrate that HCV variants reinfecting the liver graft were characterized by efficient entry and poor neutralization by antibodies present in pretransplant serum compared with variants not detected after transplantation. Monoclonal antibodies directed against HCV envelope glycoproteins or a cellular entry factor efficiently cross-neutralized infection of human hepatocytes by patient-derived viral isolates that were resistant to autologous host-neutralizing responses. These findings provide significant insights into the molecular mechanisms of viral evasion during HCV reinfection and suggest that viral entry is a viable target for prevention of HCV reinfection of the liver graft.
Figures


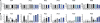

Comment in
-
Liver transplantation as a model to better understand the cell entry of hepatitis C virus.J Hepatol. 2011 Apr;54(4):825-6. doi: 10.1016/j.jhep.2010.10.036. Epub 2010 Dec 8. J Hepatol. 2011. PMID: 21167852 No abstract available.
-
Survival of the fittest: selection of hepatitis C virus variants during liver graft reinfection.Hepatology. 2011 Feb;53(2):705-8. doi: 10.1002/hep.24137. Hepatology. 2011. PMID: 21274891 No abstract available.
References
-
- Bartosch B., Verney G., Dreux M., Donot P., Morice Y., Penin F., Pawlotsky J.M., Lavillette D., Cosset F.L. 2005. An interplay between hypervariable region 1 of the hepatitis C virus E2 glycoprotein, the scavenger receptor BI, and high-density lipoprotein promotes both enhancement of infection and protection against neutralizing antibodies. J. Virol. 79:8217–8229 10.1128/JVI.79.13.8217-8229.2005 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

