Negative energy balance and hepatic gene expression patterns in high-yielding dairy cows during the early postpartum period: a global approach
- PMID: 20716645
- PMCID: PMC3008362
- DOI: 10.1152/physiolgenomics.00118.2010
Negative energy balance and hepatic gene expression patterns in high-yielding dairy cows during the early postpartum period: a global approach
Abstract
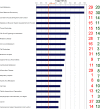
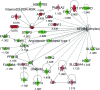
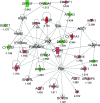
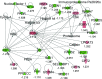
In high-yielding dairy cows the liver undergoes extensive physiological and biochemical changes during the early postpartum period in an effort to re-establish metabolic homeostasis and to counteract the adverse effects of negative energy balance (NEB). These adaptations are likely to be mediated by significant alterations in hepatic gene expression. To gain new insights into these events an energy balance model was created using differential feeding and milking regimes to produce two groups of cows with either a mild (MNEB) or severe NEB (SNEB) status. Cows were slaughtered and liver tissues collected on days 6-7 of the first follicular wave postpartum. Using an Affymetrix 23k oligonucleotide bovine array to determine global gene expression in hepatic tissue of these cows, we found a total of 416 genes (189 up- and 227 downregulated) to be altered by SNEB. Network analysis using Ingenuity Pathway Analysis revealed that SNEB was associated with widespread changes in gene expression classified into 36 gene networks including those associated with lipid metabolism, connective tissue development and function, cell signaling, cell cycle, and metabolic diseases, the three most significant of which are discussed in detail. SNEB cows displayed reduced expression of transcription activators and signal transducers that regulate the expression of genes and gene networks associated with cell signaling and tissue repair. These alterations are linked with increased expression of abnormal cell cycle and cellular proliferation associated pathways. This study provides new information and insights on the effect of SNEB on gene expression in high-yielding Holstein Friesian dairy cows in the early postpartum period.
Figures
References
-
- Bakin AV, Curran T. Role of DNA 5-methylcytosine transferase in cell transformation by fos. Science 283: 387–390, 1999. - PubMed
-
- Bernard C, Cassar-Malek I, Le Cunff M, Dubroeucq H, Renand G, Hocquette JF. New indicators of beef sensory quality revealed by expression of specific genes. J Agric Food Chem 55: 5229–5237, 2007. - PubMed
-
- Bionaz M, Drackley JK, Dann HM, Loor JJ. Liver fatty acid binding protein (FABP) and acyl-CoA synthase (ACSL) isoform gene expression due to plane of dietary energy prepartum in dairy cows. J Dairy Sci 90, Suppl 1: 972, 2007.
-
- Bionaz M, Loor JJ. Identification of reference genes for quantitative real-time PCR in the bovine mammary gland during the lactation cycle. Physiol Genomics 29: 312–319, 2007. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

