Spatial organization in bacterial chemotaxis
- PMID: 20717142
- PMCID: PMC2924652
- DOI: 10.1038/emboj.2010.178
Spatial organization in bacterial chemotaxis
Abstract
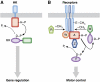
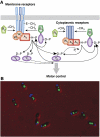
Spatial organization of signalling is not an exclusive property of eukaryotic cells. Despite the fact that bacterial signalling pathways are generally simpler than those in eukaryotes, there are several well-documented examples of higher-order intracellular signalling structures in bacteria. One of the most prominent and best-characterized structures is formed by proteins that control bacterial chemotaxis. Signals in chemotaxis are processed by ordered arrays, or clusters, of receptors and associated proteins, which amplify and integrate chemotactic stimuli in a highly cooperative manner. Receptor clusters further serve to scaffold protein interactions, enhancing the efficiency and specificity of the pathway reactions and preventing the formation of signalling gradients through the cell body. Moreover, clustering can also ensure spatial separation of multiple chemotaxis systems in one bacterium. Assembly of receptor clusters appears to be a stochastic process, but bacteria evolved mechanisms to ensure optimal cluster distribution along the cell body for partitioning to daughter cells at division.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

Comment in
-
Classifying chemoreceptors: quantity versus quality.EMBO J. 2010 Oct 20;29(20):3435-6. doi: 10.1038/emboj.2010.246. EMBO J. 2010. PMID: 20959858 Free PMC article.
References
-
- Adler J, Tso WW (1974) Decision' making in bacteria: chemotactic response of Escherichia coli to conflicting stimuli. Science 184: 1292–1294 - PubMed
-
- Alon U, Surette MG, Barkai N, Leibler S (1999) Robustness in bacterial chemotaxis. Nature 397: 168–171 - PubMed
-
- Banno S, Shiomi D, Homma M, Kawagishi I (2004) Targeting of the chemotaxis methylesterase/deamidase CheB to the polar receptor-kinase cluster in an Escherichia coli cell. Mol Microbiol 53: 1051–1063 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

