Par6 alpha interacts with the dynactin subunit p150 Glued and is a critical regulator of centrosomal protein recruitment
- PMID: 20719959
- PMCID: PMC2947473
- DOI: 10.1091/mbc.E10-05-0430
Par6 alpha interacts with the dynactin subunit p150 Glued and is a critical regulator of centrosomal protein recruitment
Abstract
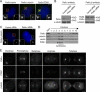
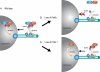
The centrosome contains proteins that control the organization of the microtubule cytoskeleton in interphase and mitosis. Its protein composition is tightly regulated through selective and cell cycle-dependent recruitment, retention, and removal of components. However, the mechanisms underlying protein delivery to the centrosome are not completely understood. We describe a novel function for the polarity protein Par6α in protein transport to the centrosome. We detected Par6α at the centrosome and centriolar satellites where it interacted with the centriolar satellite protein PCM-1 and the dynactin subunit p150(Glued). Depletion of Par6α caused the mislocalization of p150(Glued) and centrosomal components that are critical for microtubule anchoring at the centrosome. As a consequence, there were severe alterations in the organization of the microtubule cytoskeleton in the absence of Par6α and cell division was blocked. We propose a model in which Par6α controls centrosome organization through its association with the dynactin subunit p150(Glued).
Figures





Similar articles
-
Par6γ is at the mother centriole and controls centrosomal protein composition through a Par6α-dependent pathway.J Cell Sci. 2013 Feb 1;126(Pt 3):860-70. doi: 10.1242/jcs.121186. Epub 2012 Dec 21. J Cell Sci. 2013. PMID: 23264737 Free PMC article.
-
Ste20-like protein kinase SLK (LOSK) regulates microtubule organization by targeting dynactin to the centrosome.Mol Biol Cell. 2013 Oct;24(20):3205-14. doi: 10.1091/mbc.E13-03-0137. Epub 2013 Aug 28. Mol Biol Cell. 2013. PMID: 23985322 Free PMC article.
-
Cell Cycle-Dependent Localization of Dynactin Subunit p150 glued at Centrosome.J Cell Biochem. 2015 Sep;116(9):2049-60. doi: 10.1002/jcb.25160. J Cell Biochem. 2015. PMID: 25774020
-
[Dynein and dynactin as organizers of the system of cell microtubules].Ontogenez. 2006 Sep-Oct;37(5):323-39. Ontogenez. 2006. PMID: 17066975 Review. Russian.
-
Control of GABARAP-mediated autophagy by the Golgi complex, centrosome and centriolar satellites.Biol Cell. 2018 Jan;110(1):1-5. doi: 10.1111/boc.201700046. Epub 2017 Oct 27. Biol Cell. 2018. PMID: 28990689 Review.
Cited by
-
The PAR polarity complex and cerebellar granule neuron migration.Adv Exp Med Biol. 2014;800:113-31. doi: 10.1007/978-94-007-7687-6_7. Adv Exp Med Biol. 2014. PMID: 24243103 Free PMC article. Review.
-
Estrogen-responsive genes overlap with triiodothyronine-responsive genes in a breast carcinoma cell line.ScientificWorldJournal. 2014 Jan 23;2014:969404. doi: 10.1155/2014/969404. eCollection 2014. ScientificWorldJournal. 2014. PMID: 24587767 Free PMC article.
-
Inhibition of RHO-ROCK signaling enhances ICM and suppresses TE characteristics through activation of Hippo signaling in the mouse blastocyst.Dev Biol. 2014 Oct 1;394(1):142-55. doi: 10.1016/j.ydbio.2014.06.023. Epub 2014 Jul 2. Dev Biol. 2014. PMID: 24997360 Free PMC article.
-
Ccdc11 is a novel centriolar satellite protein essential for ciliogenesis and establishment of left-right asymmetry.Mol Biol Cell. 2016 Jan 1;27(1):48-63. doi: 10.1091/mbc.E15-07-0474. Epub 2015 Nov 4. Mol Biol Cell. 2016. PMID: 26538025 Free PMC article.
-
Epidermal PAR-6 and PKC-3 are essential for larval development of C. elegans and organize non-centrosomal microtubules.Elife. 2020 Dec 10;9:e62067. doi: 10.7554/eLife.62067. Elife. 2020. PMID: 33300872 Free PMC article.
References
-
- Andersen J. S., Wilkinson C. J., Mayor T., Mortensen P., Nigg E. A., Mann M. Proteomic characterization of the human centrosome by protein correlation profiling. Nature. 2003;426:570–574. - PubMed
-
- Balczon R., Varden C. E., Schroer T. A. Role for microtubules in centrosome doubling in Chinese hamster ovary cells. Cell Motil. Cytoskelet. 1999;42:60–72. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

