Telomere position effect is regulated by heterochromatin-associated proteins and NkuA in Aspergillus nidulans
- PMID: 20724388
- PMCID: PMC3068700
- DOI: 10.1099/mic.0.039255-0
Telomere position effect is regulated by heterochromatin-associated proteins and NkuA in Aspergillus nidulans
Abstract
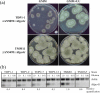
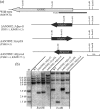
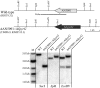
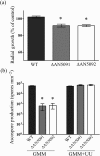
Gene-silencing mechanisms are being shown to be associated with an increasing number of fungal developmental processes. Telomere position effect (TPE) is a eukaryotic phenomenon resulting in gene repression in areas immediately adjacent to telomere caps. Here, TPE is shown to regulate expression of transgenes on the left arm of chromosome III and the right arm of chromosome VI in Aspergillus nidulans. Phenotypes found to be associated with transgene repression included reduction in radial growth and the absence of sexual spores; however, these pleiotropic phenotypes were remedied when cultures were grown on media with appropriate supplementation. Simple radial growth and ascosporogenesis assays provided insights into the mechanism of TPE, including a means to determine its extent. These experiments revealed that the KU70 homologue (NkuA) and the heterochromatin-associated proteins HepA, ClrD and HdaA were partially required for transgene silencing. This study indicates that TPE extends at least 30 kb on chromosome III, suggesting that this phenomenon may be important for gene regulation in subtelomeric regions of A. nidulans.
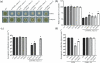
Figures


Similar articles
-
Telomerase, Ku, and telomeric silencing in Saccharomyces cerevisiae.Chromosoma. 1998 Dec;107(6-7):352-8. doi: 10.1007/s004120050318. Chromosoma. 1998. PMID: 9914366
-
A versatile and efficient gene-targeting system for Aspergillus nidulans.Genetics. 2006 Mar;172(3):1557-66. doi: 10.1534/genetics.105.052563. Epub 2005 Dec 30. Genetics. 2006. PMID: 16387870 Free PMC article.
-
Yeast Ku protein plays a direct role in telomeric silencing and counteracts inhibition by rif proteins.Curr Biol. 1999 Oct 7;9(19):1123-6. doi: 10.1016/s0960-9822(99)80483-7. Curr Biol. 1999. PMID: 10531008
-
A means to a DNA end: the many roles of Ku.Nat Rev Mol Cell Biol. 2004 May;5(5):367-78. doi: 10.1038/nrm1367. Nat Rev Mol Cell Biol. 2004. PMID: 15122350 Review. No abstract available.
-
Ethanol catabolism in Aspergillus nidulans: a model system for studying gene regulation.Prog Nucleic Acid Res Mol Biol. 2001;69:149-204. doi: 10.1016/s0079-6603(01)69047-0. Prog Nucleic Acid Res Mol Biol. 2001. PMID: 11550794 Review.
Cited by
-
Genome sequencing of evolved aspergilli populations reveals robust genomes, transversions in A. flavus, and sexual aberrancy in non-homologous end-joining mutants.BMC Biol. 2019 Nov 11;17(1):88. doi: 10.1186/s12915-019-0702-0. BMC Biol. 2019. PMID: 31711484 Free PMC article.
-
The bZIP protein MeaB mediates virulence attributes in Aspergillus flavus.PLoS One. 2013 Sep 9;8(9):e74030. doi: 10.1371/journal.pone.0074030. eCollection 2013. PLoS One. 2013. PMID: 24040154 Free PMC article.
-
A modified recombineering protocol for the genetic manipulation of gene clusters in Aspergillus fumigatus.PLoS One. 2014 Nov 5;9(11):e111875. doi: 10.1371/journal.pone.0111875. eCollection 2014. PLoS One. 2014. PMID: 25372385 Free PMC article.
-
The chromatin code of fungal secondary metabolite gene clusters.Appl Microbiol Biotechnol. 2012 Sep;95(6):1389-404. doi: 10.1007/s00253-012-4208-8. Epub 2012 Jul 20. Appl Microbiol Biotechnol. 2012. PMID: 22814413 Free PMC article. Review.
-
A novel automethylation reaction in the Aspergillus nidulans LaeA protein generates S-methylmethionine.J Biol Chem. 2013 May 17;288(20):14032-14045. doi: 10.1074/jbc.M113.465765. Epub 2013 Mar 26. J Biol Chem. 2013. PMID: 23532849 Free PMC article.
References
-
- Arnaud, M. B., Chibucos, M. C., Costanzo, M. C., Crabtree, J., Inglis, D. O., Lotia, A., Orvis, J., Shah, P., Skrzypek, M. S. & other authors (2010). The Aspergillus Genome Database, a curated comparative genomics resource for gene, protein and sequence information for the Aspergillus research community. Nucleic Acids Res 38, D420–D427. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

