The obligatory intestinal folate transporter PCFT (SLC46A1) is regulated by nuclear respiratory factor 1
- PMID: 20724482
- PMCID: PMC2962458
- DOI: 10.1074/jbc.M110.135640
The obligatory intestinal folate transporter PCFT (SLC46A1) is regulated by nuclear respiratory factor 1
Abstract
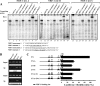
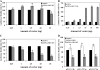
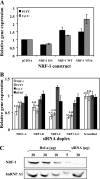
Folates are essential vitamins that play a key role as one-carbon donors in a spectrum of biosynthetic pathways including RNA and DNA synthesis. The proton-coupled folate transporter (PCFT/SLC46A1) mediates obligatory intestinal folate absorption. Loss-of-function mutations in PCFT result in hereditary folate malabsorption, an autosomal recessive disorder characterized by very low folate levels in the blood and cerebrospinal fluid. Hereditary folate malabsorption manifests within the first months after birth with anemia, immune deficiency, and neurological deficits. Here we studied the role of inducible trans-activators of PCFT gene expression. Bioinformatics identified three putative nuclear respiratory factor 1 (NRF-1) binding sites in the minimal promoter. The following evidence establish that PCFT is an NRF-1-responsive gene; electrophoretic mobility shift assay showed NRF-1 binding to native but not mutant NRF-1 sites, whereas antibody-mediated supershift analysis and chromatin immunoprecipitation revealed NRF-1 binding to its consensus sites within the PCFT promoter. Moreover, mutational inactivation of individual or all NRF-1 binding sites resulted in 40-60% decrease in luciferase reporter activity. Consistently, overexpression of NRF-1 or a constitutively active NRF-1 VP-16 construct resulted in increased reporter activity and PCFT mRNA levels. Conversely, introduction of a dominant-negative NRF-1 construct markedly repressed reporter activity and PCFT mRNA levels; likewise, introduction of NRF-1 siRNA duplexes to cells resulted in decreased PCFT transcript levels. Moreover, NRF-1 silencing down-regulated genes encoding for key folate transporters and enzymes in folate metabolism. These novel findings identify NRF-1 as a major inducible transcriptional regulator of PCFT gene expression. The implications of this linkage between folate transport and metabolism with mitochondria biogenesis and respiration are discussed.
Figures


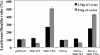
References
-
- Appling D. R. (1991) FASEB J. 5, 2645–2651 - PubMed
-
- Stockstad E. L. R. (1990) Folic Acid Metabolism in Health and Disease, pp. 1–21, Wiley-Liss, New York
-
- Assaraf Y. G. (2007) Cancer Metastasis Rev. 26, 153–181 - PubMed
-
- Dixon K. H., Lanpher B. C., Chiu J., Kelley K., Cowan K. H. (1994) J. Biol. Chem. 269, 17–20 - PubMed
-
- Matherly L. H., Hou Z., Deng Y. (2007) Cancer Metastasis Rev. 26, 111–128 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

