FLIP: a novel regulator of macrophage differentiation and granulocyte homeostasis
- PMID: 20724542
- PMCID: PMC3012591
- DOI: 10.1182/blood-2009-11-252841
FLIP: a novel regulator of macrophage differentiation and granulocyte homeostasis
Abstract
FLIP is a well-established suppressor of death receptor-mediated apoptosis. To define its essential in vivo role in myeloid cells, we generated and characterized mice with Flip conditionally deleted in the myeloid lineage. Myeloid specific Flip-deficient mice exhibited growth retardation, premature death, and splenomegaly with altered architecture and extramedullary hematopoiesis. They also displayed a dramatic increase of circulating neutrophils and multiorgan neutrophil infiltration. In contrast, although circulating inflammatory monocytes were also significantly increased, macrophages in the spleen, lymph nodes, and the peritoneal cavity were reduced. In ex vivo cultures, bone marrow progenitor cells failed to differentiate into macrophages when Flip was deleted. Mixed bone marrow chimera experiments using cells from Flip-deficient and wild-type mice did not demonstrate an inflammatory phenotype. These observations demonstrate that FLIP is necessary for macrophage differentiation and the homeostatic regulation of granulopoiesis.
Figures



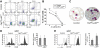



Similar articles
-
Regulation of steady-state neutrophil homeostasis by macrophages.Blood. 2011 Jan 13;117(2):618-29. doi: 10.1182/blood-2010-01-265959. Epub 2010 Oct 27. Blood. 2011. PMID: 20980680 Free PMC article.
-
Interferon regulatory factor-8-driven myeloid differentiation is regulated by 12/15-lipoxygenase-mediated redox signaling.Exp Hematol. 2010 Nov;38(11):1036-1046.e1-4. doi: 10.1016/j.exphem.2010.07.004. Epub 2010 Jul 18. Exp Hematol. 2010. PMID: 20647030 Free PMC article.
-
Mice lacking three myeloid colony-stimulating factors (G-CSF, GM-CSF, and M-CSF) still produce macrophages and granulocytes and mount an inflammatory response in a sterile model of peritonitis.J Immunol. 2007 May 15;178(10):6435-43. doi: 10.4049/jimmunol.178.10.6435. J Immunol. 2007. PMID: 17475873
-
Association of Increased F4/80high Macrophages With Suppression of Serum-Transfer Arthritis in Mice With Reduced FLIP in Myeloid Cells.Arthritis Rheumatol. 2017 Sep;69(9):1762-1771. doi: 10.1002/art.40151. Epub 2017 Aug 1. Arthritis Rheumatol. 2017. PMID: 28511285 Free PMC article.
-
Granulopoiesis and Neutrophil Homeostasis: A Metabolic, Daily Balancing Act.Trends Immunol. 2019 Jul;40(7):598-612. doi: 10.1016/j.it.2019.05.004. Trends Immunol. 2019. PMID: 31256783 Review.
Cited by
-
Myeloid-derived suppressor activity is mediated by monocytic lineages maintained by continuous inhibition of extrinsic and intrinsic death pathways.Immunity. 2014 Dec 18;41(6):947-59. doi: 10.1016/j.immuni.2014.10.020. Epub 2014 Dec 11. Immunity. 2014. PMID: 25500368 Free PMC article.
-
Mechanisms regulating the loss of Tregs in HUPO mice that develop spontaneous inflammatory arthritis.iScience. 2023 Apr 25;26(5):106734. doi: 10.1016/j.isci.2023.106734. eCollection 2023 May 19. iScience. 2023. PMID: 37216119 Free PMC article.
-
FLIP the Switch: Regulation of Apoptosis and Necroptosis by cFLIP.Int J Mol Sci. 2015 Dec 18;16(12):30321-41. doi: 10.3390/ijms161226232. Int J Mol Sci. 2015. PMID: 26694384 Free PMC article. Review.
-
Critical role of synovial tissue-resident macrophage niche in joint homeostasis and suppression of chronic inflammation.Sci Adv. 2021 Jan 6;7(2):eabd0515. doi: 10.1126/sciadv.abd0515. Print 2021 Jan. Sci Adv. 2021. PMID: 33523968 Free PMC article.
-
TAK1 control of cell death.Cell Death Differ. 2014 Nov;21(11):1667-76. doi: 10.1038/cdd.2014.123. Epub 2014 Aug 22. Cell Death Differ. 2014. PMID: 25146924 Free PMC article. Review.
References
-
- Irmler M, Thome M, Hahne M, et al. Inhibition of death receptor signals by cellular FLIP. Nature. 1997;388(6638):190–195. - PubMed
-
- Rasper DM, Vaillancourt JP, Hadano S, et al. Cell death attenuation by ‘Usurpin’, a mammalian DED-caspase homologue that precludes caspase-8 recruitment and activation by CD-95 (Fas, APO-1) receptor complex. Cell Death Diff. 1998;5(4):271–288. - PubMed
-
- Goltsev YV, Kovalenko AV, Arnold E, Varfolomeev EE, Brodianskii VM, Wallach D. CASH, a novel caspase homologue with death effector domains. J Biol Chem. 1997;272(32):19641–19644. - PubMed
-
- Micheau O, Tschopp J. Induction of TNF receptor I-mediated apoptosis via two sequential signaling complexes. Cell. 2003;114(2):181–190. - PubMed
-
- Sprick MR, Weigand MA, Rieser E, et al. FADD/MORT1 and caspase-8 are recruited to TRAIL receptors 1 and 2 and are essential for apoptosis mediated by TRAIL receptor 2. Immunity. 2000;12(6):599–609. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

