A unifying genetic model for facioscapulohumeral muscular dystrophy
- PMID: 20724583
- PMCID: PMC4677822
- DOI: 10.1126/science.1189044
A unifying genetic model for facioscapulohumeral muscular dystrophy
Abstract
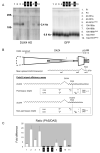
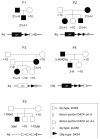
Facioscapulohumeral muscular dystrophy (FSHD) is a common form of muscular dystrophy in adults that is foremost characterized by progressive wasting of muscles in the upper body. FSHD is associated with contraction of D4Z4 macrosatellite repeats on chromosome 4q35, but this contraction is pathogenic only in certain "permissive" chromosomal backgrounds. Here, we show that FSHD patients carry specific single-nucleotide polymorphisms in the chromosomal region distal to the last D4Z4 repeat. This FSHD-predisposing configuration creates a canonical polyadenylation signal for transcripts derived from DUX4, a double homeobox gene of unknown function that straddles the last repeat unit and the adjacent sequence. Transfection studies revealed that DUX4 transcripts are efficiently polyadenylated and are more stable when expressed from permissive chromosomes. These findings suggest that FSHD arises through a toxic gain of function attributable to the stabilized distal DUX4 transcript.
Figures

Comment in
-
Genetics. Exposing a DUX tale.Science. 2010 Sep 24;329(5999):1607-8. doi: 10.1126/science.1195984. Science. 2010. PMID: 20929834 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

