DNA-Destabilizing Agents as an Alternative Approach for Targeting DNA: Mechanisms of Action and Cellular Consequences
- PMID: 20725618
- PMCID: PMC2915751
- DOI: 10.4061/2010/290935
DNA-Destabilizing Agents as an Alternative Approach for Targeting DNA: Mechanisms of Action and Cellular Consequences
Abstract
DNA targeting drugs represent a large proportion of the actual anticancer drug pharmacopeia, both in terms of drug brands and prescription volumes. Small DNA-interacting molecules share the ability of certain proteins to change the DNA helix's overall organization and geometrical orientation via tilt, roll, twist, slip, and flip effects. In this ocean of DNA-interacting compounds, most stabilize both DNA strands and very few display helix-destabilizing properties. These types of DNA-destabilizing effect are observed with certain mono- or bis-intercalators and DNA alkylating agents (some of which have been or are being developed as cancer drugs). The formation of locally destabilized DNA portions could interfere with protein/DNA recognition and potentially affect several crucial cellular processes, such as DNA repair, replication, and transcription. The present paper describes the molecular basis of DNA destabilization, the cellular impact on protein recognition, and DNA repair processes and the latter's relationships with antitumour efficacy.
Figures
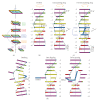
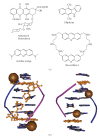
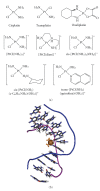


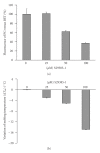
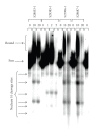
Similar articles
-
Protein Recognition in Drug-Induced DNA Alkylation: When the Moonlight Protein GAPDH Meets S23906-1/DNA Minor Groove Adducts.Int J Mol Sci. 2015 Nov 5;16(11):26555-81. doi: 10.3390/ijms161125971. Int J Mol Sci. 2015. PMID: 26556350 Free PMC article. Review.
-
Bis-benzimidazole anticancer agents: targeting human tumour helicases.Anticancer Drug Des. 1999 Feb;14(1):19-36. Anticancer Drug Des. 1999. PMID: 10363025
-
Covalent binding of antitumor benzoacronycines to double-stranded DNA induces helix opening and the formation of single-stranded DNA: unique consequences of a novel DNA-bonding mechanism.Mol Cancer Ther. 2005 Jan;4(1):71-80. Mol Cancer Ther. 2005. PMID: 15657355
-
Recent advances in small organic molecules as DNA intercalating agents: synthesis, activity, and modeling.Eur J Med Chem. 2014 Mar 3;74:95-115. doi: 10.1016/j.ejmech.2013.11.029. Epub 2014 Jan 3. Eur J Med Chem. 2014. PMID: 24448420 Review.
-
Cytotoxicity of alkylating agents towards sensitive and resistant strains of Escherichia coli in relation to extent and mode of alkylation of cellular macromolecules and repair of alkylation lesions in deoxyribonucleic acids.Biochem J. 1968 Sep;109(3):433-47. doi: 10.1042/bj1090433. Biochem J. 1968. PMID: 4879534 Free PMC article.
Cited by
-
Synthesis and Biological Evaluation of Novel 1H-Benzo[d]imidazole Derivatives as Potential Anticancer Agents Targeting Human Topoisomerase I.ACS Omega. 2022 Jan 10;7(3):2861-2880. doi: 10.1021/acsomega.1c05743. eCollection 2022 Jan 25. ACS Omega. 2022. PMID: 35097282 Free PMC article.
-
Autonomous Dynamic Control of Crown Ether Cargo Release from [2]Rotaxane Carriers in a Piperidine Oscillator.J Am Chem Soc. 2025 Jul 2;147(26):22883-22891. doi: 10.1021/jacs.5c05460. Epub 2025 Jun 17. J Am Chem Soc. 2025. PMID: 40528382 Free PMC article.
-
Disruptive DNA Intercalation Is the Mode of Interaction Behind Niacinamide Antimicrobial Activity.Microorganisms. 2025 Jul 10;13(7):1636. doi: 10.3390/microorganisms13071636. Microorganisms. 2025. PMID: 40732145 Free PMC article.
-
DNA Damage and Chromatin Conformation Changes Confer Nonhost Resistance: A Hypothesis Based on Effects of Anti-cancer Agents on Plant Defense Responses.Front Plant Sci. 2018 Jul 24;9:1056. doi: 10.3389/fpls.2018.01056. eCollection 2018. Front Plant Sci. 2018. PMID: 30087685 Free PMC article.
-
Activity of CoII-Quinalizarin: A Novel Analogue of Anthracycline-Based Anticancer Agents Targets Human DNA Topoisomerase, Whereas Quinalizarin Itself Acts via Formation of Semiquinone on Acute Lymphoblastic Leukemia MOLT-4 and HCT 116 Cells.ACS Omega. 2018 Aug 30;3(8):10255-10266. doi: 10.1021/acsomega.8b00706. eCollection 2018 Aug 31. ACS Omega. 2018. PMID: 31459155 Free PMC article.
References
-
- Gilman A, Philips FS. The biological actions and therapeutic applications of the B-chloroethyl amines and sulfides. Science. 1946;103(2675):409–436. - PubMed
-
- Watson JD, Crick FHC. Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature. 1953;171(4356):737–738. - PubMed
-
- Waring MJ. DNA modification and cancer. Annual Review of Biochemistry. 1981;50:159–192. - PubMed
-
- Teicher BA, editor. Cancer Therapeutics: Experimental and Clinical Agents (Cancer Drug Discovery and Development) [Hardcover] 1st edition. Clifton, NJ, USA: Humana Press; 1996.
-
- Braña MF, Cacho M, Gradillas A, de Pascual-Teresa B, Ramos A. Intercalators as anticancer drugs. Current Pharmaceutical Design. 2001;7(17):1745–1780. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

