Secondary structure, dynamics, and architecture of the p7 membrane protein from hepatitis C virus by NMR spectroscopy
- PMID: 20727850
- PMCID: PMC3046219
- DOI: 10.1016/j.bbamem.2010.08.010
Secondary structure, dynamics, and architecture of the p7 membrane protein from hepatitis C virus by NMR spectroscopy
Abstract
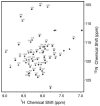
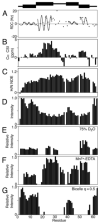
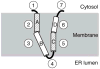
P7 is a small membrane protein that is essential for the infectivity of hepatitis C virus. Solution-state NMR experiments on p7 in DHPC micelles, including hydrogen/deuterium exchange, paramagnetic relaxation enhancement and bicelle 'q-titration,' demonstrate that the protein has a range of dynamic properties and distinct structural segments. These data along with residual dipolar couplings yield a secondary structure model of p7. We were able to confirm previous proposals that the protein has two transmembrane segments with a short interhelical loop containing the two basic residues K33 and R35. The 63-amino acid protein has a remarkably complex structure made up of seven identifiable sections, four of which are helical segments with different tilt angles and dynamics. A solid-state NMR two-dimensional separated local field spectrum of p7 aligned in phospholipid bilayers provided the tilt angles of two of these segments. A preliminary structural model of p7 derived from these NMR data is presented.
Copyright © 2010 Elsevier B.V. All rights reserved.
Figures
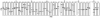

Similar articles
-
Three-dimensional structure and interaction studies of hepatitis C virus p7 in 1,2-dihexanoyl-sn-glycero-3-phosphocholine by solution nuclear magnetic resonance.Biochemistry. 2013 Aug 6;52(31):5295-303. doi: 10.1021/bi4006623. Epub 2013 Jul 24. Biochemistry. 2013. PMID: 23841474 Free PMC article.
-
The Unusual Transmembrane Partition of the Hexameric Channel of the Hepatitis C Virus.Structure. 2018 Apr 3;26(4):627-634.e4. doi: 10.1016/j.str.2018.02.011. Epub 2018 Mar 15. Structure. 2018. PMID: 29551287 Free PMC article.
-
NMR studies of p7 protein from hepatitis C virus.Eur Biophys J. 2010 Jun;39(7):1097-104. doi: 10.1007/s00249-009-0533-y. Epub 2009 Sep 2. Eur Biophys J. 2010. PMID: 19727701 Free PMC article.
-
Comparative NMR studies demonstrate profound differences between two viroporins: p7 of HCV and Vpu of HIV-1.Biochim Biophys Acta. 2011 Feb;1808(2):554-60. doi: 10.1016/j.bbamem.2010.08.005. Epub 2010 Aug 18. Biochim Biophys Acta. 2011. PMID: 20727848 Free PMC article. Review.
-
Critical Effect of the Detergent:Protein Ratio on the Formation of the Hepatitis C Virus p7 Channel.Biochemistry. 2019 Sep 17;58(37):3834-3837. doi: 10.1021/acs.biochem.9b00636. Epub 2019 Sep 3. Biochemistry. 2019. PMID: 31468972 Free PMC article. Review.
Cited by
-
Mitochondrial ADP/ATP Carrier in Dodecylphosphocholine Binds Cardiolipins with Non-native Affinity.Biophys J. 2017 Dec 5;113(11):2311-2315. doi: 10.1016/j.bpj.2017.09.019. Epub 2017 Oct 20. Biophys J. 2017. PMID: 29056231 Free PMC article.
-
Probing the orientational distribution of dyes in membranes through multiphoton microscopy.Biophys J. 2012 Sep 5;103(5):907-17. doi: 10.1016/j.bpj.2012.08.003. Biophys J. 2012. PMID: 23009840 Free PMC article.
-
Solid-State NMR for Studying the Structure and Dynamics of Viral Assemblies.Viruses. 2020 Sep 24;12(10):1069. doi: 10.3390/v12101069. Viruses. 2020. PMID: 32987909 Free PMC article. Review.
-
Targeting the Channel Activity of Viroporins.Adv Protein Chem Struct Biol. 2016;104:307-355. doi: 10.1016/bs.apcsb.2015.12.003. Epub 2016 Jan 7. Adv Protein Chem Struct Biol. 2016. PMID: 27038378 Free PMC article. Review.
-
Modeling the interaction of dodecylphosphocholine micelles with the anticoccidial peptide PW2 guided by NMR data.Molecules. 2013 Aug 20;18(8):10056-80. doi: 10.3390/molecules180810056. Molecules. 2013. PMID: 23966088 Free PMC article.
References
-
- McHutchison JG, Gordon SC, Schiff ER, Shiffman ML, Lee WM, Rustgi VK, Goodman ZD, Ling MH, Cort S, Albrecht JK. Interferon alfa-2b alone or in combination with ribavirin as initial treatment for chronic hepatitis C. Hepatitis Interventional Therapy Group. N Engl J Med. 1998;339:1485–92. - PubMed
-
- Hadziyannis SJ, Sette H, Jr, Morgan TR, Balan V, Diago M, Marcellin P, Ramadori G, Bodenheimer H, Jr, Bernstein D, Rizzetto M, Zeuzem S, Pockros PJ, Lin A, Ackrill AM. Peginterferon-alpha2a and ribavirin combination therapy in chronic hepatitis C: a randomized study of treatment duration and ribavirin dose. Ann Intern Med. 2004;140:346–55. - PubMed
-
- Di Bisceglie AM, Conjeevaram HS, Fried MW, Sallie R, Park Y, Yurdaydin C, Swain M, Kleiner DE, Mahaney K, Hoofnagle JH. Ribavirin as therapy for chronic hepatitis C. A randomized, double-blind, placebo-controlled trial. Ann Intern Med. 1995;123:897–903. - PubMed
-
- Davis GL, Esteban-Mur R, Rustgi V, Hoefs J, Gordon SC, Trepo C, Shiffman ML, Zeuzem S, Craxi A, Ling MH, Albrecht J. Interferon alfa-2b alone or in combination with ribavirin for the treatment of relapse of chronic hepatitis C. International Hepatitis Interventional Therapy Group. N Engl J Med. 1998;339:1493–9. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

