Probing tRNA interaction with biogenic polyamines
- PMID: 20729276
- PMCID: PMC2941105
- DOI: 10.1261/rna.1994310
Probing tRNA interaction with biogenic polyamines
Abstract
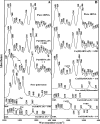
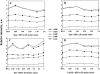
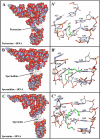
Biogenic polyamines are found to modulate protein synthesis at different levels. This effect may be explained by the ability of polyamines to bind and influence the secondary structure of tRNA, mRNA, and rRNA. We report the interaction between tRNA and the three biogenic polyamines putrescine, spermidine, spermine, and cobalt(III)hexamine at physiological conditions, using FTIR spectroscopy, capillary electrophoresis, and molecular modeling. The results indicated that tRNA was stabilized at low biogenic polyamine concentration, as a consequence of polyamine interaction with the backbone phosphate group. The main tRNA reactive sites for biogenic polyamine at low concentration were guanine-N7/O6, uracil-O2/O4, adenine-N3, and 2'OH of the ribose. At high polyamine concentration, the interaction involves guanine-N7/O6, adenine-N7, uracil-O2 reactive sites, and the backbone phosphate group. The participation of the polycation primary amino group, in the interaction and the presence of the hydrophobic contact, are also shown. The binding affinity of biogenic polyamine to tRNA molecule was in the order of spermine > spermidine > putrescine with K(Spm) = 8.7 × 10(5) M(-1), K(Spd) = 6.1 × 10(5) M(-1), and K(Put) = 1.0 × 10(5) M(-1), which correlates with their positively charged amino group content. Hill analysis showed positive cooperativity for the biogenic polyamines and negative cooperativity for cobalt-hexamine. Cobalt(III)hexamine contains high- and low-affinity sites in tRNA with K(1) = 3.2 × 10(5) M(-1) and K(2) = 1.7 × 10(5) M(-1), that have been attributed to the interactions with guanine-N7 sites and the backbone PO(2) group, respectively. This mechanism of tRNA binding could explain the condensation phenomenon observed at high Co(III) content, as previously shown in the Co(III)-DNA complexes.
Figures



Similar articles
-
Structural analysis of DNA interactions with biogenic polyamines and cobalt(III)hexamine studied by Fourier transform infrared and capillary electrophoresis.J Biol Chem. 2004 Oct 1;279(40):42041-54. doi: 10.1074/jbc.M406053200. Epub 2004 Jul 28. J Biol Chem. 2004. PMID: 15284235
-
Interaction of tRNA with antitumor polyamine analogues.Biochem Cell Biol. 2009 Aug;87(4):621-30. doi: 10.1139/o09-036. Biochem Cell Biol. 2009. PMID: 19767825
-
Binding of biogenic and synthetic polyamines to β-lactoglobulin.Int J Biol Macromol. 2011 Aug 1;49(2):201-9. doi: 10.1016/j.ijbiomac.2011.04.016. Epub 2011 May 1. Int J Biol Macromol. 2011. PMID: 21569792
-
Conjugation of biogenic and synthetic polyamines with serum proteins: A comprehensive review.Int J Biol Macromol. 2016 Nov;92:515-522. doi: 10.1016/j.ijbiomac.2016.07.049. Epub 2016 Jul 16. Int J Biol Macromol. 2016. PMID: 27431795 Review.
-
Polyamines and cellular metabolism in plants: transgenic approaches reveal different responses to diamine putrescine versus higher polyamines spermidine and spermine.Amino Acids. 2010 Feb;38(2):405-13. doi: 10.1007/s00726-009-0399-4. Epub 2009 Dec 3. Amino Acids. 2010. PMID: 19956999 Review.
Cited by
-
Binding of the 9-O-N-aryl/arylalkyl amino carbonyl methyl substituted berberine analogs to tRNA(phe.).PLoS One. 2013;8(3):e58279. doi: 10.1371/journal.pone.0058279. Epub 2013 Mar 19. PLoS One. 2013. PMID: 23526972 Free PMC article.
-
Nanoparticle-Based mRNA Vaccine Induces Protective Neutralizing Antibodies Against Infectious Bronchitis Virus in In-Vivo Infection.Vaccines (Basel). 2025 May 26;13(6):568. doi: 10.3390/vaccines13060568. Vaccines (Basel). 2025. PMID: 40573899 Free PMC article.
-
Transfer RNA Modification Enzymes from Thermophiles and Their Modified Nucleosides in tRNA.Microorganisms. 2018 Oct 20;6(4):110. doi: 10.3390/microorganisms6040110. Microorganisms. 2018. PMID: 30347855 Free PMC article. Review.
-
Clostridium thermocellum LL1210 pH homeostasis mechanisms informed by transcriptomics and metabolomics.Biotechnol Biofuels. 2018 Apr 5;11:98. doi: 10.1186/s13068-018-1095-y. eCollection 2018. Biotechnol Biofuels. 2018. PMID: 29632556 Free PMC article.
-
Conformational changes in DNA upon ligand binding monitored by circular dichroism.Int J Mol Sci. 2012;13(3):3394-3413. doi: 10.3390/ijms13033394. Epub 2012 Mar 12. Int J Mol Sci. 2012. PMID: 22489158 Free PMC article. Review.
References
-
- Agrawal RK, Penczek P, Grassucci RA, Burkhardt N, Nierhaus KH, Frank J 1999. Effect of buffer conditions on the position of tRNA on the 70 S ribosome as visualized by cryoelectron microscopy. J Biol Chem 274: 8723–8729 - PubMed
-
- Ahmed Ouameur A, Tajmir-Riahi HA 2004. Structural analysis of DNA interactions with biogenic polyamines and cobalt(III)hexamine studied by Fourier transform infrared and capillary electrophoresis. J Biol Chem 279: 42041–42054 - PubMed
-
- Ahmed Ouameur A, Malonga H, Neault J-F, Diamantoglou S, Tajmir-Riahi HA 2004a. Taxol interaction with DNA and RNA—stability and structural features. Can J Chem 82: 1112–1118
-
- Ahmed Ouameur A, Marty R, Neault J-F, Tajmir-Riahi HA 2004b. AZT binds RNA at multiple sites. DNA Cell Biol 23: 783–788 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
