Molecular basis of vancomycin dependence in VanA-type Staphylococcus aureus VRSA-9
- PMID: 20729361
- PMCID: PMC2950510
- DOI: 10.1128/JB.00613-10
Molecular basis of vancomycin dependence in VanA-type Staphylococcus aureus VRSA-9
Abstract
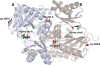
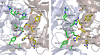
The vancomycin-resistant Staphylococcus aureus VRSA-9 clinical isolate was partially dependent on glycopeptide for growth. The responsible vanA operon had the same organization as that of Tn1546 and was located on a plasmid. The chromosomal D-Ala:D-Ala ligase (ddl) gene had two point mutations that led to Q260K and A283E substitutions, resulting in a 200-fold decrease in enzymatic activity compared to that of the wild-type strain VRSA-6. To gain insight into the mechanism of enzyme impairment, we determined the crystal structure of VRSA-9 Ddl and showed that the A283E mutation induces new ion pair/hydrogen bond interactions, leading to an asymmetric rearrangement of side chains in the dimer interface. The Q260K substitution is located in an exposed external loop and did not induce any significant conformational change. The VRSA-9 strain was susceptible to oxacillin due to synthesis of pentadepsipeptide precursors ending in D-alanyl-D-lactate which are not substrates for the β-lactam-resistant penicillin binding protein PBP2'. Comparison with the partially vancomycin-dependent VRSA-7, whose Ddl is 5-fold less efficient than that of VRSA-9, indicated that the levels of vancomycin dependence and susceptibility to β-lactams correlate with the degree of Ddl impairment. Ddl drug targeting could therefore be an effective strategy against vancomycin-resistant S. aureus.
Figures



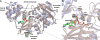
Similar articles
-
Staphylococcus aureus VRSA-11B is a constitutive vancomycin-resistant mutant of vancomycin-dependent VRSA-11A.Antimicrob Agents Chemother. 2012 Sep;56(9):4693-6. doi: 10.1128/AAC.00454-12. Epub 2012 Jun 18. Antimicrob Agents Chemother. 2012. PMID: 22710116 Free PMC article.
-
VanA-type Staphylococcus aureus strain VRSA-7 is partially dependent on vancomycin for growth.Antimicrob Agents Chemother. 2009 Sep;53(9):3657-63. doi: 10.1128/AAC.00338-09. Epub 2009 Jun 15. Antimicrob Agents Chemother. 2009. PMID: 19528271 Free PMC article.
-
The role of the Staphylococcal VraTSR regulatory system on vancomycin resistance and vanA operon expression in vancomycin-resistant Staphylococcus aureus.PLoS One. 2014 Jan 15;9(1):e85873. doi: 10.1371/journal.pone.0085873. eCollection 2014. PLoS One. 2014. PMID: 24454941 Free PMC article.
-
VanA-type vancomycin-resistant Staphylococcus aureus.Antimicrob Agents Chemother. 2009 Nov;53(11):4580-7. doi: 10.1128/AAC.00346-09. Epub 2009 Jun 8. Antimicrob Agents Chemother. 2009. PMID: 19506057 Free PMC article. Review.
-
Vancomycin Resistance in Staphylococcus aureus .Yale J Biol Med. 2017 Jun 23;90(2):269-281. eCollection 2017 Jun. Yale J Biol Med. 2017. PMID: 28656013 Free PMC article. Review.
Cited by
-
Vancomycin-resistant Staphylococcus aureus (VRSA) can overcome the cost of antibiotic resistance and may threaten vancomycin's clinical durability.PLoS Pathog. 2024 Aug 29;20(8):e1012422. doi: 10.1371/journal.ppat.1012422. eCollection 2024 Aug. PLoS Pathog. 2024. PMID: 39207957 Free PMC article.
-
Staphylococcus aureus VRSA-11B is a constitutive vancomycin-resistant mutant of vancomycin-dependent VRSA-11A.Antimicrob Agents Chemother. 2012 Sep;56(9):4693-6. doi: 10.1128/AAC.00454-12. Epub 2012 Jun 18. Antimicrob Agents Chemother. 2012. PMID: 22710116 Free PMC article.
-
VanG- and D-Ala-D-Ser-dependent peptidoglycan synthesis and vancomycin resistance in Clostridioides difficile.Mol Microbiol. 2022 Nov;118(5):526-540. doi: 10.1111/mmi.14980. Epub 2022 Sep 30. Mol Microbiol. 2022. PMID: 36065735 Free PMC article.
-
A promising approach to provide appropriate colon target drug delivery systems of vancomycin HCL: pharmaceutical and microbiological studies.Biomed Res Int. 2014;2014:182197. doi: 10.1155/2014/182197. Epub 2014 Jan 14. Biomed Res Int. 2014. PMID: 24551841 Free PMC article.
-
Fierce poison to others: the phenomenon of bacterial dependence on antibiotics.J Biomed Sci. 2023 Aug 14;30(1):67. doi: 10.1186/s12929-023-00963-x. J Biomed Sci. 2023. PMID: 37574554 Free PMC article. Review.
References
-
- Arthur, M., F. Depardieu, P. Reynolds, and P. Courvalin. 1996. Quantitative analysis of the metabolism of soluble cytoplasmic peptidoglycan precursors of glycopeptide-resistant enterococci. Mol. Microbiol. 21:33-44. - PubMed
-
- Collaborative Computational Project, Number 4. 1994. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50:760-763. - PubMed
-
- Daub, E., L. E. Zawadzke, D. Botstein, and C. T. Walsh. 1988. Isolation, cloning, and sequencing of the Salmonella typhimurium ddlA gene with purification and characterization of its product, d-alanine:d-alanine ligase (ADP forming). Biochemistry 27:3701-3708. - PubMed
-
- DeLano, W. L. 2002. The PyMol molecular graphics system. DeLano Scientific, San Carlos, CA. http://www.pymol.org.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

