Improved prediction of cardiovascular disease based on a panel of single nucleotide polymorphisms identified through genome-wide association studies
- PMID: 20729558
- PMCID: PMC3035486
- DOI: 10.1161/CIRCGENETICS.110.946269
Improved prediction of cardiovascular disease based on a panel of single nucleotide polymorphisms identified through genome-wide association studies
Abstract
Background: Genome-wide association studies (GWAS) have identified single-nucleotide polymorphisms (SNPs) at multiple loci that are significantly associated with coronary artery disease (CAD) risk. In this study, we sought to determine and compare the predictive capabilities of 9p21.3 alone and a panel of SNPs identified and replicated through GWAS for CAD.
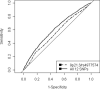
Methods and results: We used the Ottawa Heart Genomics Study (OHGS) (3323 cases, 2319 control subjects) and the Wellcome Trust Case Control Consortium (WTCCC) (1926 cases, 2938 control subjects) data sets. We compared the ability of allele counting, logistic regression, and support vector machines. Two sets of SNPs, 9p21.3 alone and a set of 12 SNPs identified by GWAS and through a model-fitting procedure, were considered. Performance was assessed by measuring area under the curve (AUC) for OHGS using 10-fold cross-validation and WTCCC as a replication set. AUC for logistic regression using OHGS increased significantly from 0.555 to 0.608 (P=3.59×10⁻¹⁴) for 9p21.3 versus the 12 SNPs, respectively. This difference remained when traditional risk factors were considered in a subgroup of OHGS (1388 cases, 2038 control subjects), with AUC increasing from 0.804 to 0.809 (P=0.037). The added predictive value over and above the traditional risk factors was not significant for 9p21.3 (AUC 0.801 versus 0.804, P=0.097) but was for the 12 SNPs (AUC 0.801 versus 0.809, P=0.0073). Performance was similar between OHGS and WTCCC. Logistic regression outperformed both support vector machines and allele counting.
Conclusions: Using the collective of 12 SNPs confers significantly greater predictive capabilities for CAD than 9p21.3, whether traditional risks are or are not considered. More accurate models probably will evolve as additional CAD-associated SNPs are identified.
Figures
References
-
- Samani NJ, Erdmann J, Hall AS, Hengstenberg C, Mangino M, Mayer B, Dixon RJ, Meitinger T, Braund P, Wichmann HE, Barrett JH, König IR, Stevens SE, Szymczak S, Tregouet DA, Iles MM, Pahlke F, Pollard H, Lieb W, Cambien F, Fischer M, Ouwehand W, Blankenberg S, Balmforth AJ, Baessler A, Ball SG, Strom TM, Braenne I, Gieger C, Deloukas P, Tobin MD, Ziegler A, Thompson JR, Schunkert H, WTCCC. the Cardiogenics Consortium Genomewide association analysis of coronary artery disease. N Engl J Med. 2007;357:443–453. - PMC - PubMed
-
- Helgadottir A, Thorleifsson G, Manolescu A, Gretarsdottir S, Blondal T, Jonasdottir A, Jonasdottir A, Sigurdsson A, Baker A, Palsson A, Masson G, Gudbjartsson DF, Magnusson KP, Andersen K, Levey AI, Backman VM, Matthiasdottir S, Jonsdottir T, Palsson S, Einarsdottir H, Gunnarsdottir S, Gylfason A, Vaccarino V, Hooper WC, Reilly MP, Granger CB, Austin H, Rader DJ, Shah SH, Quyyumi AA, Gulcher JR, Thorgeirsson G, Thorsteinsdottir U, Kong A, Stefansson K. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science. 2007;316:1491–1493. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous