Genetic dissection of intermale aggressive behavior in BALB/cJ and A/J mice
- PMID: 20731721
- PMCID: PMC3017637
- DOI: 10.1111/j.1601-183X.2010.00640.x
Genetic dissection of intermale aggressive behavior in BALB/cJ and A/J mice
Abstract
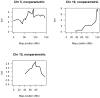
Aggressive behaviors are disabling, treatment refractory, and sometimes lethal symptoms of several neuropsychiatric disorders. However, currently available treatments for patients are inadequate, and the underlying genetics and neurobiology of aggression is only beginning to be elucidated. Inbred mouse strains are useful for identifying genomic regions, and ultimately the relevant gene variants (alleles) in these regions, that affect mammalian aggressive behaviors, which, in turn, may help to identify neurobiological pathways that mediate aggression. The BALB/cJ inbred mouse strain exhibits relatively high levels of intermale aggressive behaviors and shows multiple brain and behavioral phenotypes relevant to neuropsychiatric syndromes associated with aggression. The A/J strain shows very low levels of aggression. We hypothesized that a cross between BALB/cJ and A/J inbred strains would reveal genomic loci that influence the tendency to initiate intermale aggressive behavior. To identify such loci, we conducted a genomewide scan in an F2 population of 660 male mice bred from BALB/cJ and A/J inbred mouse strains. Three significant loci on chromosomes 5, 10 and 15 that influence aggression were identified. The chromosome 5 and 15 loci are completely novel, and the chromosome 10 locus overlaps an aggression locus mapped in our previous study that used NZB/B1NJ and A/J as progenitor strains. Haplotype analysis of BALB/cJ, NZB/B1NJ and A/J strains showed three positional candidate genes in the chromosome 10 locus. Future studies involving fine genetic mapping of these loci as well as additional candidate gene analysis may lead to an improved biological understanding of mammalian aggressive behaviors.
© 2010 The Authors. Genes, Brain and Behavior © 2010 Blackwell Publishing Ltd and International Behavioural and Neural Genetics Society.
Figures



References
-
- Appelbaum PS. Violence and mental disorders: data and public policy. American Journal of Psychiatry. 2006;163:1319–1321. - PubMed
-
- Beitchman JH, Baldassarra L, Mik H, De Luca V, King N, Bender D, Ehtesham S, Kennedy JL. Serotonin transporter polymorphisms and persistent, pervasive childhood aggression. American Journal of Psychiatry. 2006;163:1103–1105. - PubMed
-
- Boulton E, Cole C, Knight A, Cleary H, Snowden R, Plumb M. Low-penetrance genetic susceptibility and resistance loci implicated in the relative risk for radiation-induced acute myeloid leukemia in mice. Blood. 2003;101:2349–2354. - PubMed
-
- Bouwknecht JA, Paylor R. Behavioral and physiological mouse assays for anxiety: a survey in nine mouse strains. Behavioural Brain Research. 2002;136:489–501. - PubMed
-
- Brodkin ES. BALB/c mice: low sociability and other phenotypes that may be relevant to autism. Behavioural Brain Research (special issue "Animal Models for Autism") 2007;176:53–65. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

