Long-term restricted feeding alters circadian expression and reduces the level of inflammatory and disease markers
- PMID: 20731750
- PMCID: PMC4373423
- DOI: 10.1111/j.1582-4934.2010.01160.x
Long-term restricted feeding alters circadian expression and reduces the level of inflammatory and disease markers
Abstract
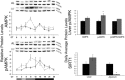
The circadian clock in peripheral tissues can be entrained by restricted feeding (RF), a regimen that restricts the duration of food availability with no calorie restriction (CR). However, it is not known whether RF can delay the occurrence of age-associated changes similar to CR. We measured circadian expression of clock genes, disease marker genes, metabolic factors and inflammatory and allergy markers in mouse serum, liver, jejunum and white adipose tissue (WAT) after long-term RF of 4 months. We found that circadian rhythmicity is more robust and is phase advanced in most of the genes and proteins tested under RF. In addition, average daily levels of some disease and inflammatory markers were reduced under RF, including liver Il-6 mRNA, tumour necrosis factor (TNF)-α and nuclear factor κB (NF-κB) protein; jejunum Arginase, Afp, Gadd45β, Il-1α and Il-1β mRNA, and interleukin (IL)-6 and TNF-α protein and WAT Il-6, Il-1β, Tnfα and Nfκb mRNA. In contrast, the anti-inflammatory cytokine Il-10 mRNA increased in the liver and jejunum. Our results suggest that RF may share some benefits with those of CR. As RF is a less harsh regimen to follow than CR, the data suggest it could be proposed for individuals seeking to improve their health.
© 2011 The Authors Journal of Cellular and Molecular Medicine © 2011 Foundation for Cellular and Molecular Medicine/Blackwell Publishing Ltd.
Figures









Similar articles
-
All-trans retinoic acid modifies the expression of clock and disease marker genes.J Nutr Biochem. 2012 Mar;23(3):209-17. doi: 10.1016/j.jnutbio.2010.11.017. Epub 2011 Apr 15. J Nutr Biochem. 2012. PMID: 21497500
-
Caffeine alters circadian rhythms and expression of disease and metabolic markers.Int J Biochem Cell Biol. 2011 May;43(5):829-38. doi: 10.1016/j.biocel.2011.02.008. Epub 2011 Feb 23. Int J Biochem Cell Biol. 2011. PMID: 21352949
-
Interleukin-1alpha stimulates proinflammatory cytokine expression in human cardiac myofibroblasts.Am J Physiol Heart Circ Physiol. 2009 Sep;297(3):H1117-27. doi: 10.1152/ajpheart.00372.2009. Epub 2009 Jul 31. Am J Physiol Heart Circ Physiol. 2009. PMID: 19648252
-
Receptor activator of nuclear factor-κB ligand is a novel inducer of myocardial inflammation.Cardiovasc Res. 2012 Apr 1;94(1):105-14. doi: 10.1093/cvr/cvs078. Epub 2012 Feb 1. Cardiovasc Res. 2012. PMID: 22298642
-
Restricted feeding entrains rhythms of inflammation-related factors without promoting an acute-phase response.Chronobiol Int. 2009 Oct;26(7):1409-29. doi: 10.3109/07420520903417003. Chronobiol Int. 2009. PMID: 19916839
Cited by
-
A fear-inducing odor alters PER2 and c-Fos expression in brain regions involved in fear memory.PLoS One. 2011;6(5):e20658. doi: 10.1371/journal.pone.0020658. Epub 2011 May 31. PLoS One. 2011. PMID: 21655193 Free PMC article.
-
Prolonged Nightly Fasting and Breast Cancer Prognosis.JAMA Oncol. 2016 Aug 1;2(8):1049-55. doi: 10.1001/jamaoncol.2016.0164. JAMA Oncol. 2016. PMID: 27032109 Free PMC article.
-
Time-Restricted Feeding Improves Body Weight Gain, Lipid Profiles, and Atherogenic Indices in Cafeteria-Diet-Fed Rats: Role of Browning of Inguinal White Adipose Tissue.Nutrients. 2020 Jul 23;12(8):2185. doi: 10.3390/nu12082185. Nutrients. 2020. PMID: 32717874 Free PMC article.
-
Obesity, cancer risk, and time-restricted eating.Cancer Metastasis Rev. 2022 Sep;41(3):697-717. doi: 10.1007/s10555-022-10061-3. Epub 2022 Aug 19. Cancer Metastasis Rev. 2022. PMID: 35984550 Free PMC article. Review.
-
A Comparison of Dietary and Caloric Restriction Models on Body Composition, Physical Performance, and Metabolic Health in Young Mice.Nutrients. 2019 Feb 7;11(2):350. doi: 10.3390/nu11020350. Nutrients. 2019. PMID: 30736418 Free PMC article.
References
-
- Lucas RJ, Freedman MS, Lupi D, et al. Identifying the photoreceptive inputs to the mammalian circadian system using transgenic and retinally degenerate mice. Behav Brain Res. 2001;125:97–102. - PubMed
-
- Lee C, Etchegaray JP, Cagampang FR, et al. Posttranslational mechanisms regulate the mammalian circadian clock. Cell. 2001;107:855–67. - PubMed
-
- Reppert SM, Weaver DR. Coordination of circadian timing in mammals. Nature. 2002;418:935–41. - PubMed
-
- Froy O, Chang DC, Reppert SM. Redox potential: differential roles in dCRY and mCRY1 functions. Curr Biol. 2002;12:147–52. - PubMed
-
- Froy O, Chapnik N, Miskin R. Long-lived alphaMUPA transgenic mice exhibit pronounced circadian rhythms. Am J Physiol Endocrinol Metab. 2006;291:E1017–24. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

