Long-lived mitochondrial (Mit) mutants of Caenorhabditis elegans utilize a novel metabolism
- PMID: 20732954
- PMCID: PMC2992375
- DOI: 10.1096/fj.10-162941
Long-lived mitochondrial (Mit) mutants of Caenorhabditis elegans utilize a novel metabolism
Abstract
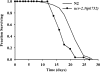
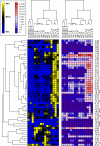
The Caenorhabditis elegans mitochondrial (Mit) mutants have disrupted mitochondrial electron transport chain (ETC) functionality, yet, surprisingly, they are long lived. We have previously proposed that Mit mutants supplement their energy needs by exploiting alternate energy production pathways normally used by wild-type animals only when exposed to hypoxic conditions. We have also proposed that longevity in the Mit mutants arises as a property of their new metabolic state. If longevity does arise as a function of metabolic state, we would expect to find a common metabolic signature among these animals. To test these predictions, we established a novel approach monitoring the C. elegans exometabolism as a surrogate marker for internal metabolic events. Using HPLC-ultraviolet-based metabolomics and multivariate analyses, we show that long-lived clk-1(qm30) and isp-1(qm150) Mit mutants have a common metabolic profile that is distinct from that of aerobically cultured wild-type animals and, unexpectedly, wild-type animals cultured under severe oxygen deprivation. Moreover, we show that 2 short-lived mitochondrial ETC mutants, mev-1(kn1) and ucr-2.3(pk732), also share a common metabolic signature that is unique. We show that removal of soluble fumarate reductase unexpectedly increases health span in several genetically defined Mit mutants, identifying at least 1 alternate energy production pathway, malate dismutation, that is operative in these animals. Our study suggests long-lived, genetically specified Mit mutants employ a novel metabolism and that life span may well arise as a function of metabolic state.
Figures
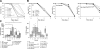

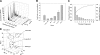
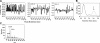
References
-
- Dillin A., Hsu A.-L., Arantes-Oliveira N., Lehrer-Graiwer J., Hsin H., Fraser A. G., Kamath R. S., Ahringer J., Kenyon C. (2002) Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401 - PubMed
-
- Lee S. S., Lee R. Y., Fraser A. G., Kamath R. S., Ahringer J., Ruvkun G. (2003) A systematic RNAi screen identifies a critical role for mitochondria in C. elegans longevity. Nat. Genet. 33, 40–48 - PubMed
-
- Rea S. L. (2005) Metabolism in the Caenorhabditis elegans Mit mutants. Exp. Gerontology 40, 841–849 - PubMed
-
- Ventura N., Rea S. L. (2007) Caenorhabditis elegans mitochondrial mutants as an investigative tool to study human neurodegenerative diseases associated with mitochondrial dysfunction. Biotechnology J. 2, 584–595 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

