Precise, facile initial rate measurements
- PMID: 20735055
- PMCID: PMC3245625
- DOI: 10.1021/jp1055528
Precise, facile initial rate measurements
Abstract
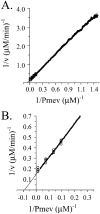
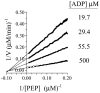
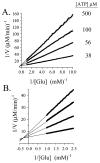
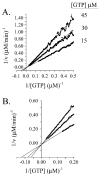
Progress curve analysis has been used sparingly in studies of enzyme-catalyzed reactions due largely to the complexity of the integrated rate expressions used in data analysis. Using an experimental design that simplifies the analysis, the advantages and limitations of progress curve experiments are explored in a study of four different enzyme-catalyzed reactions. The approach involves relatively simple protocols, requires 20-25% of the materials, and provides 10- to 20-fold signal enhancements compared to analogous initial rate studies. Product inhibition, which complicates integrated rate analysis, was circumvented using cloned, purified enzymes that remove the products and draw the reaction forward. The resulting progress curves can be transformed into the equivalent of thousands of initial rate and [S] measurements and, due to the absence of product inhibition, are plotted in the familiar, linear double-reciprocal format. Allowing product to accumulate during a reaction produces a continuously changing substrate/product ratio that can be used as the basis for obtaining product inhibition constants and to distinguish among the three classical inhibition mechanisms. Algebraic models describing the double-reciprocal patterns obtained from such inhibition studies are presented. The virtual continuum of substrate concentrations that occurs during a progress curve experiment provides a nearly errorless set of relative concentrations that results in remarkably precise data; kinetic constant standard deviations are on the order of 0.5%.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

