Zebrafish cx30.3: identification and characterization of a gap junction gene highly expressed in the skin
- PMID: 20737512
- PMCID: PMC2967642
- DOI: 10.1002/dvdy.22399
Zebrafish cx30.3: identification and characterization of a gap junction gene highly expressed in the skin
Abstract
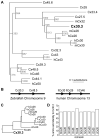
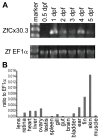
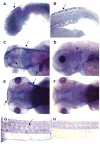
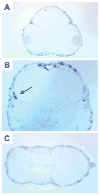
We have identified and characterized a zebrafish connexin, Cx30.3. Sequence similarity analyses suggested that Cx30.3 was orthologous to both mammalian Cx26 and Cx30, known to play important roles in the skin and inner ear of mammals. Analysis of mRNA expression showed that Cx30.3 was present in early embryos, and was highly abundant in skin, but also detected in other tissues including fins, inner ear, heart, and the retina. Injection of Cx30.3 cRNA into Xenopus oocytes elicited robust intercellular coupling with voltage gating sensitivity similar to mammalian Cx26 and Cx30. The similarities in functional properties and expression patterns suggest that Cx30.3, like mammalian Cx26 and Cx30, may play a significant role in skin development, hearing, and balance in zebrafish. Thus, zebrafish could potentially serve as an excellent model to study disorders of the skin and deafness that result from human connexin mutations.
Figures



Similar articles
-
The connexin 30.3 of zebrafish homologue of human connexin 26 may play similar role in the inner ear.Hear Res. 2014 Jul;313:55-66. doi: 10.1016/j.heares.2014.04.010. Epub 2014 May 6. Hear Res. 2014. PMID: 24811980
-
Molecular cloning, functional analysis, and RNA expression analysis of connexin45.6: a zebrafish cardiovascular connexin.Am J Physiol Heart Circ Physiol. 2004 May;286(5):H1623-32. doi: 10.1152/ajpheart.00800.2003. Epub 2004 Jan 2. Am J Physiol Heart Circ Physiol. 2004. PMID: 14704230
-
Intracellular domains of mouse connexin26 and -30 affect diffusional and electrical properties of gap junction channels.J Membr Biol. 2001 May 15;181(2):137-48. doi: 10.1007/s00232-001-0017-1. J Membr Biol. 2001. PMID: 11420600
-
Connexins and gap junctions in the inner ear--it's not just about K⁺ recycling.Cell Tissue Res. 2015 Jun;360(3):633-44. doi: 10.1007/s00441-014-2029-z. Epub 2014 Nov 9. Cell Tissue Res. 2015. PMID: 25381570 Free PMC article. Review.
-
The role of connexins in ear and skin physiology - functional insights from disease-associated mutations.Biochim Biophys Acta. 2013 Jan;1828(1):167-78. doi: 10.1016/j.bbamem.2012.06.024. Epub 2012 Jul 13. Biochim Biophys Acta. 2013. PMID: 22796187 Free PMC article. Review.
Cited by
-
Fxr signaling and microbial metabolism of bile salts in the zebrafish intestine.Sci Adv. 2021 Jul 23;7(30):eabg1371. doi: 10.1126/sciadv.abg1371. Print 2021 Jul. Sci Adv. 2021. PMID: 34301599 Free PMC article.
-
The Functional Role of CONNEXIN 26 Mutation in Nonsyndromic Hearing Loss, Demonstrated by Zebrafish Connexin 30.3 Homologue Model.Cells. 2020 May 22;9(5):1291. doi: 10.3390/cells9051291. Cells. 2020. PMID: 32455934 Free PMC article.
-
Transcriptomic Profiling of Zebrafish Hair Cells Using RiboTag.Front Cell Dev Biol. 2018 May 1;6:47. doi: 10.3389/fcell.2018.00047. eCollection 2018. Front Cell Dev Biol. 2018. PMID: 29765956 Free PMC article.
-
Single-cell RNA analysis identifies pre-migratory neural crest cells expressing markers of differentiated derivatives.Elife. 2021 Aug 16;10:e66078. doi: 10.7554/eLife.66078. Elife. 2021. PMID: 34397384 Free PMC article.
-
Connexins and pannexins in the integumentary system: the skin and appendages.Cell Mol Life Sci. 2015 Aug;72(15):2937-47. doi: 10.1007/s00018-015-1969-0. Epub 2015 Jun 20. Cell Mol Life Sci. 2015. PMID: 26091749 Free PMC article. Review.
References
-
- Beyer EC, Paul DL, Goodenough DA. Connexin family of gap junction proteins. J Membr Biol. 1990;116:187–194. - PubMed
-
- Bruzzone R, White TW, Goodenough DA. The cellular internet: On-line with connexins. Bioessays. 1996;18:709–718. - PubMed
-
- Cason N, White TW, Cheng S, Goodenough DA, Valdimarsson G. Molecular cloning, expression analysis, and functional characterization of connexin44.1: A zebrafish lens gap junction protein. Dev Dyn. 2001;221:238–247. - PubMed
-
- Cheng S, Shakespeare T, Mui R, White TW, Valdimarsson G. Connexin 48.5 is required for normal cardiovascular function and lens development in zebrafish embryos. J Biol Chem. 2004;279:36993–37003. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

