Comparative genomics and functional analysis of niche-specific adaptation in Pseudomonas putida
- PMID: 20796030
- PMCID: PMC3056050
- DOI: 10.1111/j.1574-6976.2010.00249.x
Comparative genomics and functional analysis of niche-specific adaptation in Pseudomonas putida
Abstract
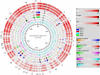
Pseudomonas putida is a gram-negative rod-shaped gammaproteobacterium that is found throughout various environments. Members of the species P. putida show a diverse spectrum of metabolic activities, which is indicative of their adaptation to various niches, which includes the ability to live in soils and sediments contaminated with high concentrations of heavy metals and organic contaminants. Pseudomonas putida strains are also found as plant growth-promoting rhizospheric and endophytic bacteria. The genome sequences of several P. putida species have become available and provide a unique tool to study the specific niche adaptation of the various P. putida strains. In this review, we compare the genomes of four P. putida strains: the rhizospheric strain KT2440, the endophytic strain W619, the aromatic hydrocarbon-degrading strain F1 and the manganese-oxidizing strain GB-1. Comparative genomics provided a powerful tool to gain new insights into the adaptation of P. putida to specific lifestyles and environmental niches, and clearly demonstrated that horizontal gene transfer played a key role in this adaptation process, as many of the niche-specific functions were found to be encoded on clearly defined genomic islands.
Journal compilation © 2010 Federation of European Microbiological Societies. Published by Blackwell Publishing Ltd. No claim to original US government works.
Figures




Similar articles
-
Insights into the genomic basis of niche specificity of Pseudomonas putida KT2440.Environ Microbiol. 2004 Dec;6(12):1264-86. doi: 10.1111/j.1462-2920.2004.00734.x. Environ Microbiol. 2004. PMID: 15560824
-
Functional genomics of the initial phase of cold adaptation of Pseudomonas putida KT2440.FEMS Microbiol Lett. 2011 May;318(1):47-54. doi: 10.1111/j.1574-6968.2011.02237.x. Epub 2011 Mar 1. FEMS Microbiol Lett. 2011. PMID: 21306427
-
Genome features of Pseudomonas putida LS46, a novel polyhydroxyalkanoate producer and its comparison with other P. putida strains.AMB Express. 2014 May 22;4:37. doi: 10.1186/s13568-014-0037-8. eCollection 2014. AMB Express. 2014. PMID: 25401060 Free PMC article.
-
Unraveling the genomic diversity of the Pseudomonas putida group: exploring taxonomy, core pangenome, and antibiotic resistance mechanisms.FEMS Microbiol Rev. 2024 Nov 23;48(6):fuae025. doi: 10.1093/femsre/fuae025. FEMS Microbiol Rev. 2024. PMID: 39390673 Free PMC article. Review.
-
Pseudomonas putida KT2440: the long journey of a soil-dweller to become a synthetic biology chassis.J Bacteriol. 2024 Jul 25;206(7):e0013624. doi: 10.1128/jb.00136-24. Epub 2024 Jul 8. J Bacteriol. 2024. PMID: 38975763 Free PMC article. Review.
Cited by
-
The complete genome sequence of the plant growth-promoting bacterium Pseudomonas sp. UW4.PLoS One. 2013;8(3):e58640. doi: 10.1371/journal.pone.0058640. Epub 2013 Mar 13. PLoS One. 2013. PMID: 23516524 Free PMC article.
-
Comparative Genomics of the Extreme Acidophile Acidithiobacillus thiooxidans Reveals Intraspecific Divergence and Niche Adaptation.Int J Mol Sci. 2016 Aug 19;17(8):1355. doi: 10.3390/ijms17081355. Int J Mol Sci. 2016. PMID: 27548157 Free PMC article.
-
Comparative genomics of Pseudomonas syringae pv. syringae strains B301D and HS191 and insights into intrapathovar traits associated with plant pathogenesis.Microbiologyopen. 2015 Aug;4(4):553-73. doi: 10.1002/mbo3.261. Epub 2015 May 4. Microbiologyopen. 2015. PMID: 25940918 Free PMC article.
-
Comparative Transcriptome Analysis of Pseudomonas putida KT2440 Revealed Its Response Mechanisms to Elevated Levels of Zinc Stress.Front Microbiol. 2018 Jul 24;9:1669. doi: 10.3389/fmicb.2018.01669. eCollection 2018. Front Microbiol. 2018. PMID: 30087671 Free PMC article.
-
Architectural Features and Resistance to Food-Grade Disinfectants in Listeria monocytogenes-Pseudomonas spp. Dual-Species Biofilms.Front Microbiol. 2022 Jun 9;13:917964. doi: 10.3389/fmicb.2022.917964. eCollection 2022. Front Microbiol. 2022. PMID: 35756028 Free PMC article.
References
-
- Alm RA, Mattick JS. Genes involved in the biogenesis and function of type-4 fimbriae in Pseudomonas aeruginosa. Gene. 1997;192:89–98. - PubMed
-
- Bingle LE, Bailey CM, Pallen MJ. Type VI secretion: a beginner's guide. Curr Opin Microbiol. 2008;11:3–8. - PubMed
-
- Braun V, Endriss F. Energy-coupled outer membrane transport proteins and regulatory proteins. Biometals. 2007;20:219–231. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

