Tobacco GTBP1, a homolog of human heterogeneous nuclear ribonucleoprotein, protects telomeres from aberrant homologous recombination
- PMID: 20798328
- PMCID: PMC2947183
- DOI: 10.1105/tpc.110.076778
Tobacco GTBP1, a homolog of human heterogeneous nuclear ribonucleoprotein, protects telomeres from aberrant homologous recombination
Abstract
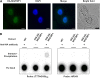
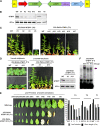
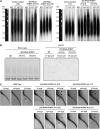
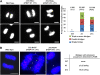
Telomeres are nucleoprotein complexes essential for the integrity of eukaryotic chromosomes. Cellular roles of single-stranded telomeric DNA binding proteins have been extensively described in yeast and animals, but our knowledge about plant single-strand telomeric factors is rudimentary. Here, we investigated Nicotiana tabacum G-strand-specific single-stranded telomere binding proteins (GTBPs), homologs of a human heterogeneous nuclear ribonucleoprotein. GTBPs bound specifically to the plant single-stranded (TTTAGGG)(4) telomeric repeat element in vitro and were associated with telomeric sequences in tobacco BY-2 suspension cells. Transgenic plants (35S:RNAi-GTBP1), in which GTBP1 was suppressed, exhibited severe developmental anomalies. In addition, the chromosomes of 35S:RNAi-GTBP1 cells displayed elongated telomeres, frequent formation of extrachromosomal telomeric circles, and numerous abnormal anaphase bridges, indicating that GTBP1 knockdown tobacco plants experienced genome instability. GTBP1 inhibited strand invasion, an initial step in interchromosomal homologous recombination. We propose that GTBP1 plays a critical role in telomere structure and function by preventing aberrant interchromosomal telomeric homologous recombination in tobacco.
Figures


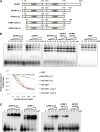
References
-
- Amiard S., et al. (2007). A topological mechanism for TRF2-enhanced strand invasion. Nat. Struct. Mol. Biol. 14: 147–154 - PubMed
-
- Bae H., Choi S.M., Yang S.W., Pai H.S., Kim W.T. (2009). Suppression of the ER-localized AAA ATPase NgCDC48 inhibits tobacco growth and development. Mol. Cells 28: 57–65 - PubMed
-
- Baumann P., Cech T.R. (2001). Pot1, the putative telomere end-binding protein in fission yeast and humans. Science 292: 1171–1175 - PubMed
-
- Bilaud T., Brun C., Ancelin K., Koering C.E., Laroche T., Gilson E. (1997). Telomeric localization of TRF2, a novel human telobox protein. Nat. Genet. 17: 236–239 - PubMed
-
- Blackburn E.H. (1991). Structure and function of telomeres. Nature 350: 569–573 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

