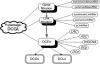
DCGL: an R package for identifying differentially coexpressed genes and links from gene expression microarray data
- PMID: 20801914
- PMCID: PMC2951087
- DOI: 10.1093/bioinformatics/btq471
DCGL: an R package for identifying differentially coexpressed genes and links from gene expression microarray data
Abstract
Summary: Gene coexpression analysis was developed to explore gene interconnection at the expression level from a systems perspective, and differential coexpression analysis (DCEA), which examines the change in gene expression correlation between two conditions, was accordingly designed as a complementary technique to traditional differential expression analysis (DEA). Since there is a shortage of DCEA tools, we implemented in an R package 'DCGL' five DCEA methods for identification of differentially coexpressed genes and differentially coexpressed links, including three currently popular methods and two novel algorithms described in a companion paper. DCGL can serve as an easy-to-use tool to facilitate differential coexpression analyses.
Contact: yyli@scbit.org and yxli@scbit.org
Supplementary information: Supplementary data are available at Bioinformatics online.
Figures
References
-
- Choi JK, et al. Differential coexpression analysis using microarray data and its application to human cancer. Bioinformatics. 2005;21:4348–4355. - PubMed
-
- Reverter A, et al. Simultaneous identification of differential gene expression and connectivity in inflammation, adipogenesis and cancer. Bioinformatics. 2006;22:2396–2404. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases