Comparative genomic characterization of Actinobacillus pleuropneumoniae
- PMID: 20802045
- PMCID: PMC2953695
- DOI: 10.1128/JB.00535-10
Comparative genomic characterization of Actinobacillus pleuropneumoniae
Abstract
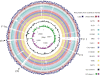
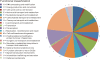
The Gram-negative bacterium Actinobacillus pleuropneumoniae is the etiologic agent of porcine contagious pleuropneumoniae, a lethal respiratory infectious disease causing great economic losses in the swine industry worldwide. In order to better interpret the genetic background of serotypic diversity, nine genomes of A. pleuropneumoniae reference strains of serovars 1, 2, 4, 6, 9, 10, 11, 12, and 13 were sequenced by using rapid high-throughput approach. Based on 12 genomes of corresponding serovar reference strains including three publicly available complete genomes (serovars 3, 5b, and 7) of this bacterium, we performed a comprehensive analysis of comparative genomics and first reported a global genomic characterization for this pathogen. Clustering of 26,012 predicted protein-coding genes showed that the pan genome of A. pleuropneumoniae consists of 3,303 gene clusters, which contain 1,709 core genome genes, 822 distributed genes, and 772 strain-specific genes. The genome components involved in the biogenesis of capsular polysaccharide and lipopolysaccharide O antigen relative to serovar diversity were compared, and their genetic diversity was depicted. Our findings shed more light on genomic features associated with serovar diversity of A. pleuropneumoniae and provide broader insight into both pathogenesis research and clinical/epidemiological application against the severe disease caused by this swine pathogen.
Figures



References
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

