Identification of a novel abscisic acid-regulated farnesol dehydrogenase from Arabidopsis
- PMID: 20807998
- PMCID: PMC2971593
- DOI: 10.1104/pp.110.157784
Identification of a novel abscisic acid-regulated farnesol dehydrogenase from Arabidopsis
Abstract
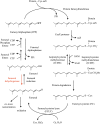
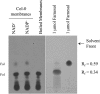
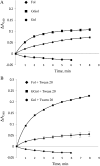
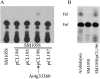
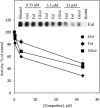
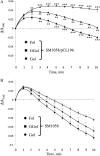
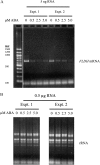
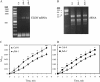
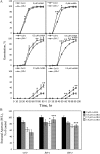
In Arabidopsis (Arabidopsis thaliana), farnesylcysteine is oxidized to farnesal and cysteine by a membrane-associated thioether oxidase called farnesylcysteine lyase. Farnesol and farnesyl phosphate kinases have also been reported in plant membranes. Together, these observations suggest the existence of enzymes that catalyze the interconversion of farnesal and farnesol. In this report, Arabidopsis membranes are shown to possess farnesol dehydrogenase activity. In addition, a gene on chromosome 4 of the Arabidopsis genome (At4g33360), called FLDH, is shown to encode an NAD(+)-dependent dehydrogenase that oxidizes farnesol more efficiently than other prenyl alcohol substrates. FLDH expression is repressed by abscisic acid (ABA) but is increased in mutants with T-DNA insertions in the FLDH 5' flanking region. These T-DNA insertion mutants, called fldh-1 and fldh-2, are associated with an ABA-insensitive phenotype, suggesting that FLDH is a negative regulator of ABA signaling.
Figures
Similar articles
-
Farnesol kinase is involved in farnesol metabolism, ABA signaling and flower development in Arabidopsis.Plant J. 2011 Jun;66(6):1078-88. doi: 10.1111/j.1365-313X.2011.04572.x. Epub 2011 Apr 12. Plant J. 2011. PMID: 21395888
-
Roles for farnesol and ABA in Arabidopsis flower development.Plant Signal Behav. 2011 Aug;6(8):1189-91. doi: 10.4161/psb.6.8.15772. Epub 2011 Aug 1. Plant Signal Behav. 2011. PMID: 21758018 Free PMC article.
-
Arabidopsis thaliana plants possess a specific farnesylcysteine lyase that is involved in detoxification and recycling of farnesylcysteine.Plant J. 2007 Jun;50(5):839-47. doi: 10.1111/j.1365-313X.2007.03091.x. Epub 2007 Apr 8. Plant J. 2007. PMID: 17425716
-
The protein phosphatase AtPP2CA negatively regulates abscisic acid signal transduction in Arabidopsis, and effects of abh1 on AtPP2CA mRNA.Plant Physiol. 2006 Jan;140(1):127-39. doi: 10.1104/pp.105.070318. Epub 2005 Dec 16. Plant Physiol. 2006. PMID: 16361522 Free PMC article.
-
The Arabidopsis A4 subfamily of lectin receptor kinases negatively regulates abscisic acid response in seed germination.Plant Physiol. 2009 Jan;149(1):434-44. doi: 10.1104/pp.108.130583. Epub 2008 Nov 5. Plant Physiol. 2009. PMID: 18987212 Free PMC article.
Cited by
-
Transcriptome analysis of Ginkgo biloba kernels.Front Plant Sci. 2015 Oct 6;6:819. doi: 10.3389/fpls.2015.00819. eCollection 2015. Front Plant Sci. 2015. PMID: 26500663 Free PMC article.
-
Physiological adventures in Candida albicans: farnesol and ubiquinones.Microbiol Mol Biol Rev. 2024 Mar 27;88(1):e0008122. doi: 10.1128/mmbr.00081-22. Epub 2024 Mar 4. Microbiol Mol Biol Rev. 2024. PMID: 38436263 Free PMC article. Review.
-
Biosynthesis of geranate via isopentenol utilization pathway in Escherichia coli.Biotechnol Bioeng. 2023 Jan;120(1):230-238. doi: 10.1002/bit.28255. Epub 2022 Nov 1. Biotechnol Bioeng. 2023. PMID: 36224741 Free PMC article.
-
Toxicity and Sublethal Effect of Farnesyl Acetate on Diamondback Moth, Plutella xylostella (L.) (Lepidoptera: Plutellidae).Insects. 2021 Jan 27;12(2):109. doi: 10.3390/insects12020109. Insects. 2021. PMID: 33513706 Free PMC article.
-
Absence of farnesol salvage in Candida albicans and probably in other fungi.Appl Environ Microbiol. 2024 Jul 24;90(7):e0087424. doi: 10.1128/aem.00874-24. Epub 2024 Jun 28. Appl Environ Microbiol. 2024. PMID: 38940563 Free PMC article.
References
-
- Alexandersson E, Saalbach G, Larsson C, Kjellbom P. (2004) Arabidopsis plasma membrane proteomics identifies components of transport, signal transduction and membrane trafficking. Plant Cell Physiol 45: 1543–1556 - PubMed
-
- Baker FC, Mauchamp B, Tsai LW, Schooley DA. (1983) Farnesol and farnesal dehydrogenase(s) in corpora allata of the tobacco hornworm moth, Manduca sexta. J Lipid Res 24: 1586–1594 - PubMed
-
- Beigneux A, Withycombe SK, Digits JA, Tschantz WR, Weinbaum CA, Griffey SM, Bergo M, Casey PJ, Young SG. (2002) Prenylcysteine lyase deficiency in mice results in the accumulation of farnesylcysteine and geranylgeranylcysteine in brain and liver. J Biol Chem 277: 38358–38363 - PubMed
-
- Bracha K, Lavy M, Yalovsky S. (2002) The Arabidopsis AtSTE24 is a CAAX protease with broad substrate specificity. J Biol Chem 277: 29856–29864 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

