Molecular and physiological analysis of drought stress in Arabidopsis reveals early responses leading to acclimation in plant growth
- PMID: 20807999
- PMCID: PMC2971604
- DOI: 10.1104/pp.110.161752
Molecular and physiological analysis of drought stress in Arabidopsis reveals early responses leading to acclimation in plant growth
Abstract
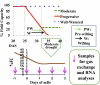
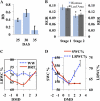
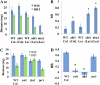
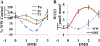
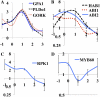
Plant drought stress response and resistance are complex biological processes that need to be analyzed at a systems level using genomics and physiological approaches to dissect experimental models that address drought stresses encountered by crops in the field. Toward this goal, a controlled, sublethal, moderate drought (mDr) treatment system was developed in Arabidopsis (Arabidopsis thaliana) as a reproducible assay for the dissection of plant responses to drought. The drought assay was validated using Arabidopsis mutants in abscisic acid (ABA) biosynthesis and signaling displaying drought sensitivity and in jasmonate response mutants showing drought resistance, indicating the crucial role of ABA and jasmonate signaling in drought response and acclimation. A comparative transcriptome analysis of soil water deficit drought stress treatments revealed the similarity of early-stage mDr to progressive drought, identifying common and specific stress-responsive genes and their promoter cis-regulatory elements. The dissection of mDr stress responses using a time-course analysis of biochemical, physiological, and molecular processes revealed early accumulation of ABA and induction of associated signaling genes, coinciding with a decrease in stomatal conductance as an early avoidance response to drought stress. This is accompanied by a peak in the expression of expansin genes involved in cell wall expansion, as a preparatory step toward drought acclimation by the adjustment of the cell wall. The time-course analysis of mDr provides a model with three stages of plant responses: an early priming and preconditioning stage, followed by an intermediate stage preparatory for acclimation, and a late stage of new homeostasis with reduced growth.
Figures




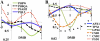


Similar articles
-
Overexpression of AtBBD1, Arabidopsis Bifunctional Nuclease, Confers Drought Tolerance by Enhancing the Expression of Regulatory Genes in ABA-Mediated Drought Stress Signaling.Int J Mol Sci. 2021 Mar 13;22(6):2936. doi: 10.3390/ijms22062936. Int J Mol Sci. 2021. PMID: 33805821 Free PMC article.
-
Functional convergence of oxylipin and abscisic acid pathways controls stomatal closure in response to drought.Plant Physiol. 2014 Mar;164(3):1151-60. doi: 10.1104/pp.113.234310. Epub 2014 Jan 15. Plant Physiol. 2014. PMID: 24429214 Free PMC article.
-
The Arabidopsis ETHYLENE RESPONSE FACTOR1 regulates abiotic stress-responsive gene expression by binding to different cis-acting elements in response to different stress signals.Plant Physiol. 2013 Jul;162(3):1566-82. doi: 10.1104/pp.113.221911. Epub 2013 May 29. Plant Physiol. 2013. PMID: 23719892 Free PMC article.
-
Mechanism of Stomatal Closure in Plants Exposed to Drought and Cold Stress.Adv Exp Med Biol. 2018;1081:215-232. doi: 10.1007/978-981-13-1244-1_12. Adv Exp Med Biol. 2018. PMID: 30288712 Review.
-
Regulatory Gene Networks in Drought Stress Responses and Resistance in Plants.Adv Exp Med Biol. 2018;1081:189-214. doi: 10.1007/978-981-13-1244-1_11. Adv Exp Med Biol. 2018. PMID: 30288711 Review.
Cited by
-
Amending clayey and sandy soils with nano - bio phosphorous for regulating tomato growth, biochemical, and physiological characteristics.Sci Rep. 2024 Oct 23;14(1):24975. doi: 10.1038/s41598-024-76389-8. Sci Rep. 2024. PMID: 39443563 Free PMC article.
-
A Dual Strategy of Breeding for Drought Tolerance and Introducing Drought-Tolerant, Underutilized Crops into Production Systems to Enhance Their Resilience to Water Deficiency.Plants (Basel). 2020 Sep 24;9(10):1263. doi: 10.3390/plants9101263. Plants (Basel). 2020. PMID: 32987964 Free PMC article. Review.
-
The Plant Cell Wall: A Complex and Dynamic Structure As Revealed by the Responses of Genes under Stress Conditions.Front Plant Sci. 2016 Aug 10;7:984. doi: 10.3389/fpls.2016.00984. eCollection 2016. Front Plant Sci. 2016. PMID: 27559336 Free PMC article. Review.
-
The agony of choice: how plants balance growth and survival under water-limiting conditions.Plant Physiol. 2013 Aug;162(4):1768-79. doi: 10.1104/pp.113.220921. Epub 2013 Jun 13. Plant Physiol. 2013. PMID: 23766368 Free PMC article. Review.
-
Analysis of melatonin regulation of germination and antioxidant metabolism in different wheat cultivars under polyethylene glycol stress.PLoS One. 2020 Aug 13;15(8):e0237536. doi: 10.1371/journal.pone.0237536. eCollection 2020. PLoS One. 2020. PMID: 32790719 Free PMC article.
References
-
- Aguirrezabal L, Bouchier-Combaud S, Radziejwoski A, Dauzat M, Cookson SJ, Granier C. (2006) Plasticity to soil water deficit in Arabidopsis thaliana: dissection of leaf development into underlying growth dynamic and cellular variables reveals invisible phenotypes. Plant Cell Environ 29: 2216–2227 - PubMed
-
- Alscher RG, Erturk N, Heath LS. (2002) Role of superoxide dismutases (SODs) in controlling oxidative stress in plants. J Exp Bot 53: 1331–1341 - PubMed
-
- Bacon M. (1999) The biochemical control of leaf expansion during drought. Plant Growth Regul 29: 101–112
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

