Computational Identification of Novel MicroRNAs and Their Targets in Vigna unguiculata
- PMID: 20811611
- PMCID: PMC2929582
- DOI: 10.1155/2010/128297
Computational Identification of Novel MicroRNAs and Their Targets in Vigna unguiculata
Abstract
MicroRNAs (miRNAs) are a class of endogenous, noncoding, short RNAs directly involved in regulating gene expression at the posttranscriptional level. High conservation of miRNAs in plant provides the foundation for identification of new miRNAs in other plant species through homology alignment. Here, previous known plant miRNAs were BLASTed against the Expressed Sequence Tag (EST) and Genomic Survey Sequence (GSS) databases of Vigna unguiculata, and according to a series of filtering criteria, a total of 47 miRNAs belonging to 13 miRNA families were identified, and 30 potential target genes of them were subsequently predicted, most of which seemed to encode transcription factors or enzymes participating in regulation of development, growth, metabolism, and other physiological processes. Overall, our findings lay the foundation for further researches of miRNAs function in Vigna unguiculata.
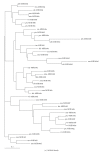
Figures











Similar articles
-
In silico identification of miRNAs and their targets from the expressed sequence tags of Raphanus sativus.Bioinformation. 2012;8(2):98-103. doi: 10.6026/97320630008098. Epub 2012 Jan 20. Bioinformation. 2012. PMID: 22359443 Free PMC article.
-
Computational identification of microRNAs and their targets in wheat (Triticum aestivum L.).Sci China C Life Sci. 2009 Nov;52(11):1091-100. doi: 10.1007/s11427-009-0144-y. Epub 2009 Nov 24. Sci China C Life Sci. 2009. PMID: 19937208
-
Bioinformatic identification and expression analysis of banana microRNAs and their targets.PLoS One. 2015 Apr 9;10(4):e0123083. doi: 10.1371/journal.pone.0123083. eCollection 2015. PLoS One. 2015. PMID: 25856313 Free PMC article.
-
Computational identification and characterization of conserved miRNAs and their target genes in garlic (Allium sativum L.) expressed sequence tags.Gene. 2014 Mar 10;537(2):333-42. doi: 10.1016/j.gene.2014.01.010. Epub 2014 Jan 13. Gene. 2014. PMID: 24434367
-
Identification and Characterization of Novel miRNAs in Chlamydomonas reinhardtii by Computational Methods.Microrna. 2016;5(1):66-77. doi: 10.2174/2211536605666160622102619. Microrna. 2016. PMID: 28105907
Cited by
-
Identification of novel soybean microRNAs involved in abiotic and biotic stresses.BMC Genomics. 2011 Jun 10;12:307. doi: 10.1186/1471-2164-12-307. BMC Genomics. 2011. PMID: 21663675 Free PMC article.
-
Functional Roles of microRNAs in Agronomically Important Plants-Potential as Targets for Crop Improvement and Protection.Front Plant Sci. 2017 Mar 22;8:378. doi: 10.3389/fpls.2017.00378. eCollection 2017. Front Plant Sci. 2017. PMID: 28382044 Free PMC article. Review.
-
Identification of miRNA encoded by Jatropha curcas from EST and GSS.Plant Signal Behav. 2013 Feb;8(2):e23152. doi: 10.4161/psb.23152. Epub 2013 Jan 8. Plant Signal Behav. 2013. PMID: 23299511 Free PMC article.
-
Identification and Characterization of MicroRNAs in Macaca fascicularis by EST Analysis.Comp Funct Genomics. 2012;2012:957607. doi: 10.1155/2012/957607. Epub 2012 Jul 5. Comp Funct Genomics. 2012. PMID: 22829752 Free PMC article.
-
Computational identification of microRNAs and their targets in cassava (Manihot esculenta Crantz.).Mol Biotechnol. 2013 Mar;53(3):257-69. doi: 10.1007/s12033-012-9521-z. Mol Biotechnol. 2013. PMID: 22388699
References
-
- Megraw M, Hatzigeorgiou AG. MicroRNA promoter analysis. Methods in Molecular Biology. 2010;592:149–161. - PubMed
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–297. - PubMed
-
- Carrington JC, Ambros V. Role of microRNAs in plant and animal development. Science. 2003;301(5631):336–338. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

