Cytokinin regulation of auxin synthesis in Arabidopsis involves a homeostatic feedback loop regulated via auxin and cytokinin signal transduction
- PMID: 20823193
- PMCID: PMC2965550
- DOI: 10.1105/tpc.110.074856
Cytokinin regulation of auxin synthesis in Arabidopsis involves a homeostatic feedback loop regulated via auxin and cytokinin signal transduction
Abstract
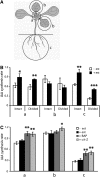
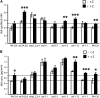
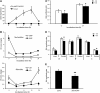
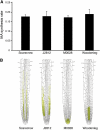
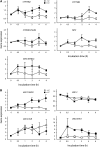
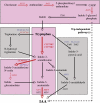
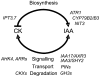
Together, auxin and cytokinin regulate many of the processes that are critical to plant growth, development, and environmental responsiveness. We have previously shown that exogenous auxin regulates cytokinin biosynthesis in Arabidopsis thaliana. In this work, we show that, conversely, the application or induced ectopic biosynthesis of cytokinin leads to a rapid increase in auxin biosynthesis in young, developing root and shoot tissues. We also show that reducing endogenous cytokinin levels, either through the induction of CYTOKININ OXIDASE expression or the mutation of one or more of the cytokinin biosynthetic ISOPENTENYLTRANSFERASE genes leads to a reduction in auxin biosynthesis. Cytokinin modifies the abundance of transcripts for several putative auxin biosynthetic genes, suggesting a direct induction of auxin biosynthesis by cytokinin. Our data indicate that cytokinin is essential, not only to maintain basal levels of auxin biosynthesis in developing root and shoot tissues but also for the dynamic regulation of auxin biosynthesis in response to changing developmental or environmental conditions. In combination with our previous work, the data suggest that a homeostatic feedback regulatory loop involving both auxin and cytokinin signaling acts to maintain appropriate auxin and cytokinin concentrations in developing root and shoot tissues.
Figures
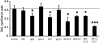
References
-
- Baker D.A. (2000). Long-distance vascular transport of endogenous hormones in plants and their role in source:sink regulation. Isr. J. Plant Sci. 48: 199–203
-
- Birnbaum K., Jung J.W., Wang J.Y., Lambert G.M., Hirst J.A., Galbraith D.W., Benfey P.N. (2005). Cell type-specific expression profiling in plants via cell sorting of protoplasts from fluorescent reporter lines. Nat. Methods 2: 615–619 - PubMed
-
- Casimiro I., Beeckman T., Graham N., Bhalerao R., Zhang H., Casero P., Sandberg G., Bennett M.J. (2003). Dissecting Arabidopsis lateral root development. Trends Plant Sci. 8: 165–171 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
- BB/E01772X/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BBS/B/1356X/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/I001271/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/E022758/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- G17764/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

